Research report guide: Definition, types, and tips
Last updated
5 March 2024
Reviewed by
Short on time? Get an AI generated summary of this article instead
From successful product launches or software releases to planning major business decisions, research reports serve many vital functions. They can summarize evidence and deliver insights and recommendations to save companies time and resources. They can reveal the most value-adding actions a company should take.
However, poorly constructed reports can have the opposite effect! Taking the time to learn established research-reporting rules and approaches will equip you with in-demand skills. You’ll be able to capture and communicate information applicable to numerous situations and industries, adding another string to your resume bow.
- What are research reports?
A research report is a collection of contextual data, gathered through organized research, that provides new insights into a particular challenge (which, for this article, is business-related). Research reports are a time-tested method for distilling large amounts of data into a narrow band of focus.
Their effectiveness often hinges on whether the report provides:
Strong, well-researched evidence
Comprehensive analysis
Well-considered conclusions and recommendations
Though the topic possibilities are endless, an effective research report keeps a laser-like focus on the specific questions or objectives the researcher believes are key to achieving success. Many research reports begin as research proposals, which usually include the need for a report to capture the findings of the study and recommend a course of action.
A description of the research method used, e.g., qualitative, quantitative, or other
Statistical analysis
Causal (or explanatory) research (i.e., research identifying relationships between two variables)
Inductive research, also known as ‘theory-building’
Deductive research, such as that used to test theories
Action research, where the research is actively used to drive change
- Importance of a research report
Research reports can unify and direct a company's focus toward the most appropriate strategic action. Of course, spending resources on a report takes up some of the company's human and financial resources. Choosing when a report is called for is a matter of judgment and experience.
Some development models used heavily in the engineering world, such as Waterfall development, are notorious for over-relying on research reports. With Waterfall development, there is a linear progression through each step of a project, and each stage is precisely documented and reported on before moving to the next.
The pace of the business world is faster than the speed at which your authors can produce and disseminate reports. So how do companies strike the right balance between creating and acting on research reports?
The answer lies, again, in the report's defined objectives. By paring down your most pressing interests and those of your stakeholders, your research and reporting skills will be the lenses that keep your company's priorities in constant focus.
Honing your company's primary objectives can save significant amounts of time and align research and reporting efforts with ever-greater precision.
Some examples of well-designed research objectives are:
Proving whether or not a product or service meets customer expectations
Demonstrating the value of a service, product, or business process to your stakeholders and investors
Improving business decision-making when faced with a lack of time or other constraints
Clarifying the relationship between a critical cause and effect for problematic business processes
Prioritizing the development of a backlog of products or product features
Comparing business or production strategies
Evaluating past decisions and predicting future outcomes
- Features of a research report
Research reports generally require a research design phase, where the report author(s) determine the most important elements the report must contain.
Just as there are various kinds of research, there are many types of reports.
Here are the standard elements of almost any research-reporting format:
Report summary. A broad but comprehensive overview of what readers will learn in the full report. Summaries are usually no more than one or two paragraphs and address all key elements of the report. Think of the key takeaways your primary stakeholders will want to know if they don’t have time to read the full document.
Introduction. Include a brief background of the topic, the type of research, and the research sample. Consider the primary goal of the report, who is most affected, and how far along the company is in meeting its objectives.
Methods. A description of how the researcher carried out data collection, analysis, and final interpretations of the data. Include the reasons for choosing a particular method. The methods section should strike a balance between clearly presenting the approach taken to gather data and discussing how it is designed to achieve the report's objectives.
Data analysis. This section contains interpretations that lead readers through the results relevant to the report's thesis. If there were unexpected results, include here a discussion on why that might be. Charts, calculations, statistics, and other supporting information also belong here (or, if lengthy, as an appendix). This should be the most detailed section of the research report, with references for further study. Present the information in a logical order, whether chronologically or in order of importance to the report's objectives.
Conclusion. This should be written with sound reasoning, often containing useful recommendations. The conclusion must be backed by a continuous thread of logic throughout the report.
- How to write a research paper
With a clear outline and robust pool of research, a research paper can start to write itself, but what's a good way to start a research report?
Research report examples are often the quickest way to gain inspiration for your report. Look for the types of research reports most relevant to your industry and consider which makes the most sense for your data and goals.
The research report outline will help you organize the elements of your report. One of the most time-tested report outlines is the IMRaD structure:
Introduction
...and Discussion
Pay close attention to the most well-established research reporting format in your industry, and consider your tone and language from your audience's perspective. Learn the key terms inside and out; incorrect jargon could easily harm the perceived authority of your research paper.
Along with a foundation in high-quality research and razor-sharp analysis, the most effective research reports will also demonstrate well-developed:
Internal logic
Narrative flow
Conclusions and recommendations
Readability, striking a balance between simple phrasing and technical insight

How to gather research data for your report
The validity of research data is critical. Because the research phase usually occurs well before the writing phase, you normally have plenty of time to vet your data.
However, research reports could involve ongoing research, where report authors (sometimes the researchers themselves) write portions of the report alongside ongoing research.
One such research-report example would be an R&D department that knows its primary stakeholders are eager to learn about a lengthy work in progress and any potentially important outcomes.
However you choose to manage the research and reporting, your data must meet robust quality standards before you can rely on it. Vet any research with the following questions in mind:
Does it use statistically valid analysis methods?
Do the researchers clearly explain their research, analysis, and sampling methods?
Did the researchers provide any caveats or advice on how to interpret their data?
Have you gathered the data yourself or were you in close contact with those who did?
Is the source biased?
Usually, flawed research methods become more apparent the further you get through a research report.
It's perfectly natural for good research to raise new questions, but the reader should have no uncertainty about what the data represents. There should be no doubt about matters such as:
Whether the sampling or analysis methods were based on sound and consistent logic
What the research samples are and where they came from
The accuracy of any statistical functions or equations
Validation of testing and measuring processes
When does a report require design validation?
A robust design validation process is often a gold standard in highly technical research reports. Design validation ensures the objects of a study are measured accurately, which lends more weight to your report and makes it valuable to more specialized industries.
Product development and engineering projects are the most common research-report examples that typically involve a design validation process. Depending on the scope and complexity of your research, you might face additional steps to validate your data and research procedures.
If you’re including design validation in the report (or report proposal), explain and justify your data-collection processes. Good design validation builds greater trust in a research report and lends more weight to its conclusions.
Choosing the right analysis method
Just as the quality of your report depends on properly validated research, a useful conclusion requires the most contextually relevant analysis method. This means comparing different statistical methods and choosing the one that makes the most sense for your research.
Most broadly, research analysis comes down to quantitative or qualitative methods (respectively: measurable by a number vs subjectively qualified values). There are also mixed research methods, which bridge the need for merging hard data with qualified assessments and still reach a cohesive set of conclusions.
Some of the most common analysis methods in research reports include:
Significance testing (aka hypothesis analysis), which compares test and control groups to determine how likely the data was the result of random chance.
Regression analysis , to establish relationships between variables, control for extraneous variables , and support correlation analysis.
Correlation analysis (aka bivariate testing), a method to identify and determine the strength of linear relationships between variables. It’s effective for detecting patterns from complex data, but care must be exercised to not confuse correlation with causation.
With any analysis method, it's important to justify which method you chose in the report. You should also provide estimates of the statistical accuracy (e.g., the p-value or confidence level of quantifiable data) of any data analysis.
This requires a commitment to the report's primary aim. For instance, this may be achieving a certain level of customer satisfaction by analyzing the cause and effect of changes to how service is delivered. Even better, use statistical analysis to calculate which change is most positively correlated with improved levels of customer satisfaction.
- Tips for writing research reports
There's endless good advice for writing effective research reports, and it almost all depends on the subjective aims of the people behind the report. Due to the wide variety of research reports, the best tips will be unique to each author's purpose.
Consider the following research report tips in any order, and take note of the ones most relevant to you:
No matter how in depth or detailed your report might be, provide a well-considered, succinct summary. At the very least, give your readers a quick and effective way to get up to speed.
Pare down your target audience (e.g., other researchers, employees, laypersons, etc.), and adjust your voice for their background knowledge and interest levels
For all but the most open-ended research, clarify your objectives, both for yourself and within the report.
Leverage your team members’ talents to fill in any knowledge gaps you might have. Your team is only as good as the sum of its parts.
Justify why your research proposal’s topic will endure long enough to derive value from the finished report.
Consolidate all research and analysis functions onto a single user-friendly platform. There's no reason to settle for less than developer-grade tools suitable for non-developers.
What's the format of a research report?
The research-reporting format is how the report is structured—a framework the authors use to organize their data, conclusions, arguments, and recommendations. The format heavily determines how the report's outline develops, because the format dictates the overall structure and order of information (based on the report's goals and research objectives).
What's the purpose of a research-report outline?
A good report outline gives form and substance to the report's objectives, presenting the results in a readable, engaging way. For any research-report format, the outline should create momentum along a chain of logic that builds up to a conclusion or interpretation.
What's the difference between a research essay and a research report?
There are several key differences between research reports and essays:
Research report:
Ordered into separate sections
More commercial in nature
Often includes infographics
Heavily descriptive
More self-referential
Usually provides recommendations
Research essay
Does not rely on research report formatting
More academically minded
Normally text-only
Less detailed
Omits discussion of methods
Usually non-prescriptive
Should you be using a customer insights hub?
Do you want to discover previous research faster?
Do you share your research findings with others?
Do you analyze research data?
Start for free today, add your research, and get to key insights faster
Editor’s picks
Last updated: 18 April 2023
Last updated: 27 February 2023
Last updated: 6 February 2023
Last updated: 5 February 2023
Last updated: 16 April 2023
Last updated: 9 March 2023
Last updated: 30 April 2024
Last updated: 12 December 2023
Last updated: 11 March 2024
Last updated: 4 July 2024
Last updated: 6 March 2024
Last updated: 5 March 2024
Last updated: 13 May 2024
Latest articles
Related topics, .css-je19u9{-webkit-align-items:flex-end;-webkit-box-align:flex-end;-ms-flex-align:flex-end;align-items:flex-end;display:-webkit-box;display:-webkit-flex;display:-ms-flexbox;display:flex;-webkit-flex-direction:row;-ms-flex-direction:row;flex-direction:row;-webkit-box-flex-wrap:wrap;-webkit-flex-wrap:wrap;-ms-flex-wrap:wrap;flex-wrap:wrap;-webkit-box-pack:center;-ms-flex-pack:center;-webkit-justify-content:center;justify-content:center;row-gap:0;text-align:center;max-width:671px;}@media (max-width: 1079px){.css-je19u9{max-width:400px;}.css-je19u9>span{white-space:pre;}}@media (max-width: 799px){.css-je19u9{max-width:400px;}.css-je19u9>span{white-space:pre;}} decide what to .css-1kiodld{max-height:56px;display:-webkit-box;display:-webkit-flex;display:-ms-flexbox;display:flex;-webkit-align-items:center;-webkit-box-align:center;-ms-flex-align:center;align-items:center;}@media (max-width: 1079px){.css-1kiodld{display:none;}} build next, decide what to build next, log in or sign up.
Get started for free
- Research Report: Definition, Types + [Writing Guide]

One of the reasons for carrying out research is to add to the existing body of knowledge. Therefore, when conducting research, you need to document your processes and findings in a research report.
With a research report, it is easy to outline the findings of your systematic investigation and any gaps needing further inquiry. Knowing how to create a detailed research report will prove useful when you need to conduct research.
What is a Research Report?
A research report is a well-crafted document that outlines the processes, data, and findings of a systematic investigation. It is an important document that serves as a first-hand account of the research process, and it is typically considered an objective and accurate source of information.
In many ways, a research report can be considered as a summary of the research process that clearly highlights findings, recommendations, and other important details. Reading a well-written research report should provide you with all the information you need about the core areas of the research process.
Features of a Research Report
So how do you recognize a research report when you see one? Here are some of the basic features that define a research report.
- It is a detailed presentation of research processes and findings, and it usually includes tables and graphs.
- It is written in a formal language.
- A research report is usually written in the third person.
- It is informative and based on first-hand verifiable information.
- It is formally structured with headings, sections, and bullet points.
- It always includes recommendations for future actions.
Types of Research Report
The research report is classified based on two things; nature of research and target audience.
Nature of Research
- Qualitative Research Report
This is the type of report written for qualitative research . It outlines the methods, processes, and findings of a qualitative method of systematic investigation. In educational research, a qualitative research report provides an opportunity for one to apply his or her knowledge and develop skills in planning and executing qualitative research projects.
A qualitative research report is usually descriptive in nature. Hence, in addition to presenting details of the research process, you must also create a descriptive narrative of the information.
- Quantitative Research Report
A quantitative research report is a type of research report that is written for quantitative research. Quantitative research is a type of systematic investigation that pays attention to numerical or statistical values in a bid to find answers to research questions.
In this type of research report, the researcher presents quantitative data to support the research process and findings. Unlike a qualitative research report that is mainly descriptive, a quantitative research report works with numbers; that is, it is numerical in nature.
Target Audience
Also, a research report can be said to be technical or popular based on the target audience. If you’re dealing with a general audience, you would need to present a popular research report, and if you’re dealing with a specialized audience, you would submit a technical report.
- Technical Research Report
A technical research report is a detailed document that you present after carrying out industry-based research. This report is highly specialized because it provides information for a technical audience; that is, individuals with above-average knowledge in the field of study.
In a technical research report, the researcher is expected to provide specific information about the research process, including statistical analyses and sampling methods. Also, the use of language is highly specialized and filled with jargon.
Examples of technical research reports include legal and medical research reports.
- Popular Research Report
A popular research report is one for a general audience; that is, for individuals who do not necessarily have any knowledge in the field of study. A popular research report aims to make information accessible to everyone.
It is written in very simple language, which makes it easy to understand the findings and recommendations. Examples of popular research reports are the information contained in newspapers and magazines.
Importance of a Research Report
- Knowledge Transfer: As already stated above, one of the reasons for carrying out research is to contribute to the existing body of knowledge, and this is made possible with a research report. A research report serves as a means to effectively communicate the findings of a systematic investigation to all and sundry.
- Identification of Knowledge Gaps: With a research report, you’d be able to identify knowledge gaps for further inquiry. A research report shows what has been done while hinting at other areas needing systematic investigation.
- In market research, a research report would help you understand the market needs and peculiarities at a glance.
- A research report allows you to present information in a precise and concise manner.
- It is time-efficient and practical because, in a research report, you do not have to spend time detailing the findings of your research work in person. You can easily send out the report via email and have stakeholders look at it.
Guide to Writing a Research Report
A lot of detail goes into writing a research report, and getting familiar with the different requirements would help you create the ideal research report. A research report is usually broken down into multiple sections, which allows for a concise presentation of information.
Structure and Example of a Research Report
This is the title of your systematic investigation. Your title should be concise and point to the aims, objectives, and findings of a research report.
- Table of Contents
This is like a compass that makes it easier for readers to navigate the research report.
An abstract is an overview that highlights all important aspects of the research including the research method, data collection process, and research findings. Think of an abstract as a summary of your research report that presents pertinent information in a concise manner.
An abstract is always brief; typically 100-150 words and goes straight to the point. The focus of your research abstract should be the 5Ws and 1H format – What, Where, Why, When, Who and How.
- Introduction
Here, the researcher highlights the aims and objectives of the systematic investigation as well as the problem which the systematic investigation sets out to solve. When writing the report introduction, it is also essential to indicate whether the purposes of the research were achieved or would require more work.
In the introduction section, the researcher specifies the research problem and also outlines the significance of the systematic investigation. Also, the researcher is expected to outline any jargons and terminologies that are contained in the research.
- Literature Review
A literature review is a written survey of existing knowledge in the field of study. In other words, it is the section where you provide an overview and analysis of different research works that are relevant to your systematic investigation.
It highlights existing research knowledge and areas needing further investigation, which your research has sought to fill. At this stage, you can also hint at your research hypothesis and its possible implications for the existing body of knowledge in your field of study.
- An Account of Investigation
This is a detailed account of the research process, including the methodology, sample, and research subjects. Here, you are expected to provide in-depth information on the research process including the data collection and analysis procedures.
In a quantitative research report, you’d need to provide information surveys, questionnaires and other quantitative data collection methods used in your research. In a qualitative research report, you are expected to describe the qualitative data collection methods used in your research including interviews and focus groups.
In this section, you are expected to present the results of the systematic investigation.
This section further explains the findings of the research, earlier outlined. Here, you are expected to present a justification for each outcome and show whether the results are in line with your hypotheses or if other research studies have come up with similar results.
- Conclusions
This is a summary of all the information in the report. It also outlines the significance of the entire study.
- References and Appendices
This section contains a list of all the primary and secondary research sources.
Tips for Writing a Research Report
- Define the Context for the Report
As is obtainable when writing an essay, defining the context for your research report would help you create a detailed yet concise document. This is why you need to create an outline before writing so that you do not miss out on anything.
- Define your Audience
Writing with your audience in mind is essential as it determines the tone of the report. If you’re writing for a general audience, you would want to present the information in a simple and relatable manner. For a specialized audience, you would need to make use of technical and field-specific terms.
- Include Significant Findings
The idea of a research report is to present some sort of abridged version of your systematic investigation. In your report, you should exclude irrelevant information while highlighting only important data and findings.
- Include Illustrations
Your research report should include illustrations and other visual representations of your data. Graphs, pie charts, and relevant images lend additional credibility to your systematic investigation.
- Choose the Right Title
A good research report title is brief, precise, and contains keywords from your research. It should provide a clear idea of your systematic investigation so that readers can grasp the entire focus of your research from the title.
- Proofread the Report
Before publishing the document, ensure that you give it a second look to authenticate the information. If you can, get someone else to go through the report, too, and you can also run it through proofreading and editing software.
How to Gather Research Data for Your Report
- Understand the Problem
Every research aims at solving a specific problem or set of problems, and this should be at the back of your mind when writing your research report. Understanding the problem would help you to filter the information you have and include only important data in your report.
- Know what your report seeks to achieve
This is somewhat similar to the point above because, in some way, the aim of your research report is intertwined with the objectives of your systematic investigation. Identifying the primary purpose of writing a research report would help you to identify and present the required information accordingly.
- Identify your audience
Knowing your target audience plays a crucial role in data collection for a research report. If your research report is specifically for an organization, you would want to present industry-specific information or show how the research findings are relevant to the work that the company does.
- Create Surveys/Questionnaires
A survey is a research method that is used to gather data from a specific group of people through a set of questions. It can be either quantitative or qualitative.
A survey is usually made up of structured questions, and it can be administered online or offline. However, an online survey is a more effective method of research data collection because it helps you save time and gather data with ease.
You can seamlessly create an online questionnaire for your research on Formplus . With the multiple sharing options available in the builder, you would be able to administer your survey to respondents in little or no time.
Formplus also has a report summary too l that you can use to create custom visual reports for your research.
Step-by-step guide on how to create an online questionnaire using Formplus
- Sign into Formplus
In the Formplus builder, you can easily create different online questionnaires for your research by dragging and dropping preferred fields into your form. To access the Formplus builder, you will need to create an account on Formplus.
Once you do this, sign in to your account and click on Create new form to begin.
- Edit Form Title : Click on the field provided to input your form title, for example, “Research Questionnaire.”
- Edit Form : Click on the edit icon to edit the form.
- Add Fields : Drag and drop preferred form fields into your form in the Formplus builder inputs column. There are several field input options for questionnaires in the Formplus builder.
- Edit fields
- Click on “Save”
- Form Customization: With the form customization options in the form builder, you can easily change the outlook of your form and make it more unique and personalized. Formplus allows you to change your form theme, add background images, and even change the font according to your needs.
- Multiple Sharing Options: Formplus offers various form-sharing options, which enables you to share your questionnaire with respondents easily. You can use the direct social media sharing buttons to share your form link to your organization’s social media pages. You can also send out your survey form as email invitations to your research subjects too. If you wish, you can share your form’s QR code or embed it on your organization’s website for easy access.
Conclusion
Always remember that a research report is just as important as the actual systematic investigation because it plays a vital role in communicating research findings to everyone else. This is why you must take care to create a concise document summarizing the process of conducting any research.
In this article, we’ve outlined essential tips to help you create a research report. When writing your report, you should always have the audience at the back of your mind, as this would set the tone for the document.

Connect to Formplus, Get Started Now - It's Free!
- ethnographic research survey
- research report
- research report survey
- busayo.longe

You may also like:
Assessment Tools: Types, Examples & Importance
In this article, you’ll learn about different assessment tools to help you evaluate performance in various contexts

How to Write a Problem Statement for your Research
Learn how to write problem statements before commencing any research effort. Learn about its structure and explore examples
21 Chrome Extensions for Academic Researchers in 2022
In this article, we will discuss a number of chrome extensions you can use to make your research process even seamless
Ethnographic Research: Types, Methods + [Question Examples]
Simple guide on ethnographic research, it types, methods, examples and advantages. Also highlights how to conduct an ethnographic...
Formplus - For Seamless Data Collection
Collect data the right way with a versatile data collection tool. try formplus and transform your work productivity today..
- Academic Skills
- Reading, writing and referencing
Research reports
This resource will help you identify the common elements and basic format of a research report.
Research reports generally follow a similar structure and have common elements, each with a particular purpose. Learn more about each of these elements below.
Common elements of reports
Your title should be brief, topic-specific, and informative, clearly indicating the purpose and scope of your study. Include key words in your title so that search engines can easily access your work. For example: Measurement of water around Station Pier.
An abstract is a concise summary that helps readers to quickly assess the content and direction of your paper. It should be brief, written in a single paragraph and cover: the scope and purpose of your report; an overview of methodology; a summary of the main findings or results; principal conclusions or significance of the findings; and recommendations made.
The information in the abstract must be presented in the same order as it is in your report. The abstract is usually written last when you have developed your arguments and synthesised the results.
The introduction creates the context for your research. It should provide sufficient background to allow the reader to understand and evaluate your study without needing to refer to previous publications. After reading the introduction your reader should understand exactly what your research is about, what you plan to do, why you are undertaking this research and which methods you have used. Introductions generally include:
- The rationale for the present study. Why are you interested in this topic? Why is this topic worth investigating?
- Key terms and definitions.
- An outline of the research questions and hypotheses; the assumptions or propositions that your research will test.
Not all research reports have a separate literature review section. In shorter research reports, the review is usually part of the Introduction.
A literature review is a critical survey of recent relevant research in a particular field. The review should be a selection of carefully organised, focused and relevant literature that develops a narrative ‘story’ about your topic. Your review should answer key questions about the literature:
- What is the current state of knowledge on the topic?
- What differences in approaches / methodologies are there?
- Where are the strengths and weaknesses of the research?
- What further research is needed? The review may identify a gap in the literature which provides a rationale for your study and supports your research questions and methodology.
The review is not just a summary of all you have read. Rather, it must develop an argument or a point of view that supports your chosen methodology and research questions.
The purpose of this section is to detail how you conducted your research so that others can understand and replicate your approach.
You need to briefly describe the subjects (if appropriate), any equipment or materials used and the approach taken. If the research method or method of data analysis is commonly used within your field of study, then simply reference the procedure. If, however, your methods are new or controversial then you need to describe them in more detail and provide a rationale for your approach. The methodology is written in the past tense and should be as concise as possible.
This section is a concise, factual summary of your findings, listed under headings appropriate to your research questions. It’s common to use tables and graphics. Raw data or details about the method of statistical analysis used should be included in the Appendices.
Present your results in a consistent manner. For example, if you present the first group of results as percentages, it will be confusing for the reader and difficult to make comparisons of data if later results are presented as fractions or as decimal values.
In general, you won’t discuss your results here. Any analysis of your results usually occurs in the Discussion section.
Notes on visual data representation:
- Graphs and tables may be used to reveal trends in your data, but they must be explained and referred to in adjacent accompanying text.
- Figures and tables do not simply repeat information given in the text: they summarise, amplify or complement it.
- Graphs are always referred to as ‘Figures’, and both axes must be clearly labelled.
- Tables must be numbered, and they must be able to stand-alone or make sense without your reader needing to read all of the accompanying text.
The Discussion responds to the hypothesis or research question. This section is where you interpret your results, account for your findings and explain their significance within the context of other research. Consider the adequacy of your sampling techniques, the scope and long-term implications of your study, any problems with data collection or analysis and any assumptions on which your study was based. This is also the place to discuss any disappointing results and address limitations.
Checklist for the discussion
- To what extent was each hypothesis supported?
- To what extent are your findings validated or supported by other research?
- Were there unexpected variables that affected your results?
- On reflection, was your research method appropriate?
- Can you account for any differences between your results and other studies?
Conclusions in research reports are generally fairly short and should follow on naturally from points raised in the Discussion. In this section you should discuss the significance of your findings. To what extent and in what ways are your findings useful or conclusive? Is further research required? If so, based on your research experience, what suggestions could you make about improvements to the scope or methodology of future studies?
Also, consider the practical implications of your results and any recommendations you could make. For example, if your research is on reading strategies in the primary school classroom, what are the implications of your results for the classroom teacher? What recommendations could you make for teachers?
A Reference List contains all the resources you have cited in your work, while a Bibliography is a wider list containing all the resources you have consulted (but not necessarily cited) in the preparation of your work. It is important to check which of these is required, and the preferred format, style of references and presentation requirements of your own department.
Appendices (singular ‘Appendix’) provide supporting material to your project. Examples of such materials include:
- Relevant letters to participants and organisations (e.g. regarding the ethics or conduct of the project).
- Background reports.
- Detailed calculations.
Different types of data are presented in separate appendices. Each appendix must be titled, labelled with a number or letter, and referred to in the body of the report.
Appendices are placed at the end of a report, and the contents are generally not included in the word count.
Fi nal ti p
While there are many common elements to research reports, it’s always best to double check the exact requirements for your task. You may find that you don’t need some sections, can combine others or have specific requirements about referencing, formatting or word limits.

Looking for one-on-one advice?
Get tailored advice from an Academic Skills Adviser by booking an Individual appointment, or get quick feedback from one of our Academic Writing Mentors via email through our Writing advice service.
Go to Student appointments
- Skip to main content
- Skip to primary sidebar
- Skip to footer
- QuestionPro

- Solutions Industries Gaming Automotive Sports and events Education Government Travel & Hospitality Financial Services Healthcare Cannabis Technology Use Case AskWhy Communities Audience Contactless surveys Mobile LivePolls Member Experience GDPR Positive People Science 360 Feedback Surveys
- Resources Blog eBooks Survey Templates Case Studies Training Help center
Home Market Research
Research Reports: Definition and How to Write Them

Reports are usually spread across a vast horizon of topics but are focused on communicating information about a particular topic and a niche target market. The primary motive of research reports is to convey integral details about a study for marketers to consider while designing new strategies.
Certain events, facts, and other information based on incidents need to be relayed to the people in charge, and creating research reports is the most effective communication tool. Ideal research reports are extremely accurate in the offered information with a clear objective and conclusion. These reports should have a clean and structured format to relay information effectively.
What are Research Reports?
Research reports are recorded data prepared by researchers or statisticians after analyzing the information gathered by conducting organized research, typically in the form of surveys or qualitative methods .
A research report is a reliable source to recount details about a conducted research. It is most often considered to be a true testimony of all the work done to garner specificities of research.
The various sections of a research report are:
- Background/Introduction
- Implemented Methods
- Results based on Analysis
- Deliberation
Learn more: Quantitative Research
Components of Research Reports
Research is imperative for launching a new product/service or a new feature. The markets today are extremely volatile and competitive due to new entrants every day who may or may not provide effective products. An organization needs to make the right decisions at the right time to be relevant in such a market with updated products that suffice customer demands.
The details of a research report may change with the purpose of research but the main components of a report will remain constant. The research approach of the market researcher also influences the style of writing reports. Here are seven main components of a productive research report:
- Research Report Summary: The entire objective along with the overview of research are to be included in a summary which is a couple of paragraphs in length. All the multiple components of the research are explained in brief under the report summary. It should be interesting enough to capture all the key elements of the report.
- Research Introduction: There always is a primary goal that the researcher is trying to achieve through a report. In the introduction section, he/she can cover answers related to this goal and establish a thesis which will be included to strive and answer it in detail. This section should answer an integral question: “What is the current situation of the goal?”. After the research design was conducted, did the organization conclude the goal successfully or they are still a work in progress – provide such details in the introduction part of the research report.
- Research Methodology: This is the most important section of the report where all the important information lies. The readers can gain data for the topic along with analyzing the quality of provided content and the research can also be approved by other market researchers . Thus, this section needs to be highly informative with each aspect of research discussed in detail. Information needs to be expressed in chronological order according to its priority and importance. Researchers should include references in case they gained information from existing techniques.
- Research Results: A short description of the results along with calculations conducted to achieve the goal will form this section of results. Usually, the exposition after data analysis is carried out in the discussion part of the report.
Learn more: Quantitative Data
- Research Discussion: The results are discussed in extreme detail in this section along with a comparative analysis of reports that could probably exist in the same domain. Any abnormality uncovered during research will be deliberated in the discussion section. While writing research reports, the researcher will have to connect the dots on how the results will be applicable in the real world.
- Research References and Conclusion: Conclude all the research findings along with mentioning each and every author, article or any content piece from where references were taken.
Learn more: Qualitative Observation
15 Tips for Writing Research Reports
Writing research reports in the manner can lead to all the efforts going down the drain. Here are 15 tips for writing impactful research reports:
- Prepare the context before starting to write and start from the basics: This was always taught to us in school – be well-prepared before taking a plunge into new topics. The order of survey questions might not be the ideal or most effective order for writing research reports. The idea is to start with a broader topic and work towards a more specific one and focus on a conclusion or support, which a research should support with the facts. The most difficult thing to do in reporting, without a doubt is to start. Start with the title, the introduction, then document the first discoveries and continue from that. Once the marketers have the information well documented, they can write a general conclusion.
- Keep the target audience in mind while selecting a format that is clear, logical and obvious to them: Will the research reports be presented to decision makers or other researchers? What are the general perceptions around that topic? This requires more care and diligence. A researcher will need a significant amount of information to start writing the research report. Be consistent with the wording, the numbering of the annexes and so on. Follow the approved format of the company for the delivery of research reports and demonstrate the integrity of the project with the objectives of the company.
- Have a clear research objective: A researcher should read the entire proposal again, and make sure that the data they provide contributes to the objectives that were raised from the beginning. Remember that speculations are for conversations, not for research reports, if a researcher speculates, they directly question their own research.
- Establish a working model: Each study must have an internal logic, which will have to be established in the report and in the evidence. The researchers’ worst nightmare is to be required to write research reports and realize that key questions were not included.
Learn more: Quantitative Observation
- Gather all the information about the research topic. Who are the competitors of our customers? Talk to other researchers who have studied the subject of research, know the language of the industry. Misuse of the terms can discourage the readers of research reports from reading further.
- Read aloud while writing. While reading the report, if the researcher hears something inappropriate, for example, if they stumble over the words when reading them, surely the reader will too. If the researcher can’t put an idea in a single sentence, then it is very long and they must change it so that the idea is clear to everyone.
- Check grammar and spelling. Without a doubt, good practices help to understand the report. Use verbs in the present tense. Consider using the present tense, which makes the results sound more immediate. Find new words and other ways of saying things. Have fun with the language whenever possible.
- Discuss only the discoveries that are significant. If some data are not really significant, do not mention them. Remember that not everything is truly important or essential within research reports.
Learn more: Qualitative Data
- Try and stick to the survey questions. For example, do not say that the people surveyed “were worried” about an research issue , when there are different degrees of concern.
- The graphs must be clear enough so that they understand themselves. Do not let graphs lead the reader to make mistakes: give them a title, include the indications, the size of the sample, and the correct wording of the question.
- Be clear with messages. A researcher should always write every section of the report with an accuracy of details and language.
- Be creative with titles – Particularly in segmentation studies choose names “that give life to research”. Such names can survive for a long time after the initial investigation.
- Create an effective conclusion: The conclusion in the research reports is the most difficult to write, but it is an incredible opportunity to excel. Make a precise summary. Sometimes it helps to start the conclusion with something specific, then it describes the most important part of the study, and finally, it provides the implications of the conclusions.
- Get a couple more pair of eyes to read the report. Writers have trouble detecting their own mistakes. But they are responsible for what is presented. Ensure it has been approved by colleagues or friends before sending the find draft out.
Learn more: Market Research and Analysis
MORE LIKE THIS

Life@QuestionPro: Thomas Maiwald-Immer’s Experience
Aug 9, 2024

Top 13 Reporting Tools to Transform Your Data Insights & More
Aug 8, 2024

Employee Satisfaction: How to Boost Your Workplace Happiness?
Aug 7, 2024

Jotform vs Formstack: Which Form Builder Should You Choose?
Other categories.
- Academic Research
- Artificial Intelligence
- Assessments
- Brand Awareness
- Case Studies
- Communities
- Consumer Insights
- Customer effort score
- Customer Engagement
- Customer Experience
- Customer Loyalty
- Customer Research
- Customer Satisfaction
- Employee Benefits
- Employee Engagement
- Employee Retention
- Friday Five
- General Data Protection Regulation
- Insights Hub
- Life@QuestionPro
- Market Research
- Mobile diaries
- Mobile Surveys
- New Features
- Online Communities
- Question Types
- Questionnaire
- QuestionPro Products
- Release Notes
- Research Tools and Apps
- Revenue at Risk
- Survey Templates
- Training Tips
- Tuesday CX Thoughts (TCXT)
- Uncategorized
- What’s Coming Up
- Workforce Intelligence
Want to create or adapt books like this? Learn more about how Pressbooks supports open publishing practices.
Section 1- Evidence-based practice (EBP)
Chapter 6: Components of a Research Report
Components of a research report.
Partido, B.B.
Elements of research report
| Introduction | What is the issue? |
| Methods | What methods have been used to investigate the issue? |
| Results | What was found? |
| Discussion | What are the implications of the findings? |
The research report contains four main areas:
- Introduction – What is the issue? What is known? What is not known? What are you trying to find out? This sections ends with the purpose and specific aims of the study.
- Methods – The recipe for the study. If someone wanted to perform the same study, what information would they need? How will you answer your research question? This part usually contains subheadings: Participants, Instruments, Procedures, Data Analysis,
- Results – What was found? This is organized by specific aims and provides the results of the statistical analysis.
- Discussion – How do the results fit in with the existing literature? What were the limitations and areas of future research?
Formalized Curiosity for Knowledge and Innovation Copyright © by partido1. All Rights Reserved.
Uncomplicated Reviews of Educational Research Methods
- Writing a Research Report
.pdf version of this page
This review covers the basic elements of a research report. This is a general guide for what you will see in journal articles or dissertations. This format assumes a mixed methods study, but you can leave out either quantitative or qualitative sections if you only used a single methodology.
This review is divided into sections for easy reference. There are five MAJOR parts of a Research Report:
1. Introduction 2. Review of Literature 3. Methods 4. Results 5. Discussion
As a general guide, the Introduction, Review of Literature, and Methods should be about 1/3 of your paper, Discussion 1/3, then Results 1/3.
Section 1 : Cover Sheet (APA format cover sheet) optional, if required.
Section 2: Abstract (a basic summary of the report, including sample, treatment, design, results, and implications) (≤ 150 words) optional, if required.
Section 3 : Introduction (1-3 paragraphs) • Basic introduction • Supportive statistics (can be from periodicals) • Statement of Purpose • Statement of Significance
Section 4 : Research question(s) or hypotheses • An overall research question (optional) • A quantitative-based (hypotheses) • A qualitative-based (research questions) Note: You will generally have more than one, especially if using hypotheses.
Section 5: Review of Literature ▪ Should be organized by subheadings ▪ Should adequately support your study using supporting, related, and/or refuting evidence ▪ Is a synthesis, not a collection of individual summaries
Section 6: Methods ▪ Procedure: Describe data gathering or participant recruitment, including IRB approval ▪ Sample: Describe the sample or dataset, including basic demographics ▪ Setting: Describe the setting, if applicable (generally only in qualitative designs) ▪ Treatment: If applicable, describe, in detail, how you implemented the treatment ▪ Instrument: Describe, in detail, how you implemented the instrument; Describe the reliability and validity associated with the instrument ▪ Data Analysis: Describe type of procedure (t-test, interviews, etc.) and software (if used)
Section 7: Results ▪ Restate Research Question 1 (Quantitative) ▪ Describe results ▪ Restate Research Question 2 (Qualitative) ▪ Describe results
Section 8: Discussion ▪ Restate Overall Research Question ▪ Describe how the results, when taken together, answer the overall question ▪ ***Describe how the results confirm or contrast the literature you reviewed
Section 9: Recommendations (if applicable, generally related to practice)
Section 10: Limitations ▪ Discuss, in several sentences, the limitations of this study. ▪ Research Design (overall, then info about the limitations of each separately) ▪ Sample ▪ Instrument/s ▪ Other limitations
Section 11: Conclusion (A brief closing summary)
Section 12: References (APA format)
Share this:
About research rundowns.
Research Rundowns was made possible by support from the Dewar College of Education at Valdosta State University .
- Experimental Design
- What is Educational Research?
- Writing Research Questions
- Mixed Methods Research Designs
- Qualitative Coding & Analysis
- Qualitative Research Design
- Correlation
- Effect Size
- Instrument, Validity, Reliability
- Mean & Standard Deviation
- Significance Testing (t-tests)
- Steps 1-4: Finding Research
- Steps 5-6: Analyzing & Organizing
- Steps 7-9: Citing & Writing
Create a free website or blog at WordPress.com.

- Already have a WordPress.com account? Log in now.
- Subscribe Subscribed
- Copy shortlink
- Report this content
- View post in Reader
- Manage subscriptions
- Collapse this bar

Research Report Meaning, Characteristics and Types
Table of contents:-, research report meaning, characteristics of good research report, key characteristics of research report, types of research report, stages in preparation of research report, characteristics of a good report.
A research report is a document that conveys the outcomes of a study or investigation. Its purpose is to communicate the research’s findings, conclusions, and implications to a particular audience. This report aims to offer a comprehensive and unbiased overview of the research process, methodology, and results.
Once the researcher has completed data collection , data processing, developing and testing hypotheses, and interpretation of responses, the next important phase in research is the preparation of the research report. A research report is essential for the communication of research findings to its potential users.
The research report must be free from personal bias, external influences, and subjective factors. i.e., it must be free from one’s liking and disliking. The research report must be prepared to meet impersonal needs.
What is Research Report?
According to Lancaster, “A report is a statement of collected and considered facts, so drawn-ups to give clear and concise information to persons who are not already in possession of the full facts of the subject matter of the report”.
When researchers communicate their results in writing, they create a research report. It includes the research methodology, approaches, data collection precautions, research findings, and recommendations for solving related problems. Managers can put this result into action for more effective decision making .
Generally, top management places a higher emphasis on obtaining the research outcome rather than delving into the research procedure. Hence, the research report acts as a presentation that highlights the procedure and methodology adopted by the researcher.
The research report presents the complete procedure in a comprehensive way that in turn helps the management in making crucial decisions. Creating a research report adheres to a specific format, sequence, and writing style.
Enhance the effectiveness of a research report by incorporating various charts, graphs, diagrams, tables, etc. By using different representation techniques, researchers can convince the audience as well as the management in an effective way.
Characteristics of a good research report are listed below:
- Clarity and Completeness
- Reliability
- Comprehensibility and Readability
- Logical Content

The following paragraphs outline the characteristics of a good research report.
1) Accuracy
Report information must be accurate and based on facts, credible sources and data to establish reliability and trustworthiness. It should not be biased by the personal feelings of the writer. The information presented must be as precise as possible.
2) Simplicity
The language of a research report should be as simple as possible to ensure easy understanding. A good report communicates its message clearly and without ambiguity through its language.
It is a document of practical utility; therefore, it should be grammatically accurate, brief, and easily understood.
Jargon and technical words should be avoided when writing the report. Even in a technical report, there should be restricted use of technical terms if it is to be presented to laymen.
3) Clarity and Completeness
The report must be straightforward, lucid, and comprehensive in every aspect. Ambiguity should be avoided at all costs. Clarity is achieved through the strategic and practical organization of information. Report writers should divide their report into short paragraphs with headings and insert other suitable signposts to enhance clarity. They should:
- Approach their task systematically,
- Clarify their purpose,
- Define their sources,
- State their findings and
- Make necessary recommendations.
A report should concisely convey the key points without unnecessary length, ensuring that the reader’s patience is not lost and ideas are not confused. Many times, people lack the time to read lengthy reports.
However, a report must also be complete. Sometimes, it is important to have a detailed discussion about the facts. A report is not an essay; therefore, points should be added to it.
5) Appearance
A report requires a visually appealing presentation and, whenever feasible, should be attention-grabbing. An effective report depends on the arrangement, organization, format, layout, typography, printing quality, and paper choice. Big companies often produce very attractive and colourful Annual Reports to showcase their achievements and financial performance.
6) Comprehensibility and Readability
Reports should be clear and straightforward for easy understanding. The style of presentation and the choice of words should be attractive to readers. The writer must present the facts in elegant and grammatically correct English so that the reader is compelled to read the report from beginning to end.
Only then does a report serve its purpose. A report written by different individuals on the same subject matter can vary depending on the intended audience.
7) Reliability
Reports should be reliable and should not create an erroneous impression in the minds of readers due to oversight or neglect. The facts presented in a report should be pertinent.
Every fact in a report must align with the central purpose, but it is also vital to ensure that all pertinent information is included.
Irrelevant facts can make a report confusing, and the exclusion of relevant facts can render it incomplete and likely to mislead.
Report writing should not incur unnecessary expenses. Cost-effective methods should be used to maintain a consistent level of quality when communicating the content.
9) Timelines
Reports can be valuable and practical when they reach the readers promptly. Any delay in the submission of reports renders the preparation of reports futile and sometimes obsolete.
10) Logical Content
The points mentioned in a report should be arranged in a step-by-step logical sequence and not haphazardly. Distinctive points should have self-explanatory headings and sub-headings. The scientific accuracy of facts is very essential for a report.
Planning is necessary before a report is prepared, as reports invariably lead to decision-making, and inaccurate facts may result in unsuccessful decisions.
Related Articles:
- nature of marketing
- difference between questionnaire and schedule
- features of marginal costing
- placement in hrm
- limitations of marginal costing
- nature of leadership
- difference between advertising and personal selling
A research report serves as a means of communicating research findings to the readers effectively.

- Clarity in Information
- Optimal Length
- Objective and Simple Language
- Clear Thinking and Logical Organization
- Engaging Style
- Clarity in Presentation
- Readability
- Best Composition Practices
- Inferences and Conclusions
- Proper References
- Attractive Appearance
i) Clarity in Information
A well-defined research report must define the what, why, who, whom, when, where, and how of the research study. It must help the readers to understand the focus of the information presented.
ii) Optimal Length
The report should strike a balance, being sufficiently brief and appropriately extended. It should cover the subject matter adequately while maintaining the reader’s interest.
iii) Objective and Simple Language
The report should be written in an objective style, employing simple language. Correctness, precision, and clarity should be prioritized, avoiding wordiness, indirection, and pompous language.
iv) Clear Thinking and Logical Organization
An excellent report integrates clear thinking, logical organization, and sound interpretation of the research findings.
v) Engaging Style
It should not be dull; instead, it should captivate and sustain the reader’s interest.
vi) Accuracy
Accuracy is paramount. The report must present facts objectively, eschewing exaggerations and superlatives.
vii) Clarity in Presentation
Presentation clarity is achieved through familiar words, unambiguous statements, and explicit definitions of new concepts or terms.
viii) Coherence
The logical flow of ideas and a coherent sequence of sentences contribute to a smooth continuity of thought.
ix) Readability
Even technical reports should be easily understandable. Translate technicalities into reader-friendly language.
x) Best Composition Practices
Follow best composition practices, ensuring readability through proper paragraphing, short sentences, and the use of illustrations, examples, section headings, charts, graphs, and diagrams.
xi) Inferences and Conclusions
Draw sound inferences and conclusions from statistical tables without repeating them in verbal form.
xii) Proper References
Footnote references should be correctly formatted, and the bibliography should be reasonably complete.
xiii) Attractive Appearance
The report should be visually appealing, maintaining a neat and clean appearance, whether typed or printed.
xiv) Error-Free
The report should be free from all types of mistakes, including language, factual, spelling, and calculation errors.
In striving for these qualities, the researcher enhances the overall quality of the report.
Research reports are of the following types:
- Technical Report
- Manuscripts for Journal Articles
- Thesis and Dissertations
- Other Types of Research Report

1) Technical Report
Technical reports are reports which contain detailed information about the research problem and its findings. These reports are typically subject to review by individuals interested in research methodology. Such reports include detailed descriptions of used methods for research design such as universe selection , sample preparation, designing questionnaire , identifying potential data sources, etc. These reports provide a complete description of every step, method, and tool used. When crafting technical reports, we assume that users possess knowledge of research methodology, which is why the language used in these reports is technical. Technical reports are valuable in situations where there is a need for statistical analysis of collected data. Researchers also employ it in conducting a series of research studies, where they can repetitively use the methodology.
2) Manuscripts for Journal Articles
When authors prepare a report with a particular layout or design for publishing in an academic or scientific journal, it becomes a “manuscript for journal articles”. Journal articles are a concise and complete presentation of a particular research study. While technical reports present a detailed description of all the activities in research, journal articles are known for presenting only a few critical areas or findings of a study. The readers or audience of journal articles include other researchers, management and executives, strategic analysts and the general public, interested in the topic.
In general, a manuscript for a journal article typically ranges from 10 to 30 pages in length. Sometimes there is a page or word limit for preparing the report. Authors primarily submit manuscripts for journal articles online, although they occasionally send paper copies through regular mail.
3) Thesis and Dissertations
Students working towards a Master’s, PhD, or another higher degree generally produce a thesis or dissertation, which is a form of research report. Like other normal research reports, the thesis or dissertation usually describes the design, tools or methods and results of the student’s research in detail.
These reports typically include a detailed section called the literature review, which encompasses relevant literature and previous studies on the topic. Firstly, the work or research of the student is analysed by a professional researcher or an expert in that particular research field, and then the thesis is written under the guidance of a professional supervisor. Dissertations and theses usually span approximately 120 to 300 pages in length.
Generally, the university or institution decides the length of the dissertation or thesis. A distinctive feature of a thesis or a dissertation is that it is quite economical, as it requires few printed and bound copies of the report. Sometimes electronic copies are required to be submitted along with the hard copy of the thesis or dissertations. Compact discs (CDs) are used to generate the electronic copy.
4) Other Types of Research Report
Along with the above-mentioned types, there are some other types of research reports, which are as follows:
- Popular Report
- Interim Report
- Summary Report
- Research Abstract
i) Popular Report
A popular report is prepared for the use of administrators, executives, or managers. It is simple and attractive in the form of a report. Clear and concise statements are used with less technical or statistical terms. Data representation is kept very simple through minimal use of graphs and charts. It has a different format than that of a technical one by liberally using margins and blank spaces. The style of writing a popular report is journalistic and precise. It is written to facilitate reading rapidly and comprehending quickly.
ii) Interim Report
An interim report is a kind of report which is prepared to show the sponsors, the progress of research work before the final presentation of the report. It is prepared when there is a certain time gap between the data collection and presentation. In this scenario, the completed portion of data analysis along with its findings is described in a particular interim report.
iii) Summary Report
This type of report is related to the interest of the general public. The findings of such a report are helpful for the decision making of general users. The language used for preparing a summary report is comprehensive and simple. The inclusion of numerous graphs and tables enhances the report’s overall clarity and comprehension. The main focus of this report is on the objectives, findings, and implications of the research issue.
iv) Research Abstract
The research abstract is a short presentation of the technical report. All the elements of a particular technical report, such as the research problem, objectives, sampling techniques, etc., are described in the research abstract but the description is concise and easy.
Research reports result from meticulous and deliberate work. Consequently, the preparation of the information can be delineated into the following key stages:
1) Logical Understanding and Subject Analysis: This stage involves a comprehensive grasp and analysis of the subject matter.
2) Planning/Designing the Final Outline: In this phase, the final outline of the report is meticulously planned and designed.
3) Write-Up/Preparation of Rough Draft: The report takes shape during this stage through the composition of a rough draft.
4) Polishing/Finalization of the Research Report: The final stage encompasses refining and polishing the report to achieve its ultimate form.
(adsbygoogle = window.adsbygoogle || []).push({});
Logical understanding and subject analysis.
This initial stage focuses on the subject’s development, which can be achieved through two approaches:
- Logical development and
- Chronological development
Logical development relies on mental connections and associations between different aspects facilitated by rational analysis. Typically, this involves progressing from simple to complex elements. In contrast, chronological development follows a sequence of time or events, with instructions or descriptions often adhering to chronological order.
Designing the Final Outline of the Research Report
This marks the second stage in report writing. Once the subject matter is comprehended, the subsequent step involves structuring the report, arranging its components, and outlining them. This stage is also referred to as the planning and organization stage. While ideas may flow through the author’s mind, they must create a plan, sketch, or design. These are necessary for achieving a harmonious succession to become more accessible, and the author may be unsure where to commence or conclude. Effective communication of research results hinges not only on language but predominantly on the meticulous planning and organization of the report.
Preparation of the Rough Draft
The third stage involves the writing and drafting of the report. This phase is pivotal for the researcher as they translate their research study into written form, articulating what they have accomplished and how they intend to convey it.
The clarity in communication and reporting during this stage is influenced by several factors, including the audience, the technical complexity of the problem, the researcher’s grasp of facts and techniques, their proficiency in the language (communication skills), the completeness of notes and documentation, and the availability of analyzed results.
Depending on these factors, some authors may produce the report with just one or two drafts. In contrast, others, with less command over language and a lack of clarity about the problem and subject matter, may require more time and multiple drafts (first draft, second draft, third draft, fourth draft, etc.).
Finalization of the Research Report
This marks the last stage, potentially the most challenging phase in all formal writing. Constructing the structure is relatively easy, but refining and adding the finishing touches require considerable time. Consider, for instance, the construction of a house. The work progresses swiftly up to the roofing (structure) stage, but the final touches and completion demand a significant amount of time.
The rough draft, whether it is the second draft or the n th draft, must undergo rewriting and polishing to meet the requirements. The meticulous revision of the rough draft is what distinguishes a mediocre piece of writing from a good one. During the polishing and finalization phase, it is crucial to scrutinize the report for weaknesses in the logical development of the subject and the cohesion of its presentation. Additionally, attention should be given to the mechanics of writing, including language, usage, grammar, spelling, and punctuation.
Good research possesses certain characteristics, which are as follows:
- Empirical Basis
- Logical Approach
- Systematic Nature
- Replicability
- Validity and Verifiability
- Theory and Principle Development
1. Empirical Basis: It implies that any conclusion drawn is grounded in hardcore evidence collected from real-life experiences and observations. This foundation provides external validity to research results.
2. Logical Approach: Good research is logical, guided by the rules of reasoning and analytical processes of induction (general to specific) and deduction (particular to the public). Logical reasoning is integral to making research feasible and meaningful in decision-making.
3. Systematic Nature: Good research is systematic, which adheres to a structured set of rules, following specific steps in a defined sequence. Systematic research encourages creative thinking while avoiding reliance on guesswork and intuition to reach conclusions.
4. Replicability: Scientific research designs, procedures, and results should be replicable. This ensures that anyone apart from the original researcher can assess their validity. Researchers can use or replicate results obtained by others, making the procedures and outcomes of the research both replicable and transmittable.
5. Validity and Verifiability: Good research involves precise observation and accurate description. The researcher selects reliable and valid instruments for data collection, employing statistical measures to portray results accurately. The conclusions drawn are correct and verifiable by both the researcher and others.
6. Theory and Principle Development: It contributes to formulating theories and principles, aiding accurate predictions about the variables under study. By making sound generalizations based on observed samples, researchers extend their findings beyond immediate situations, objects, or groups, formulating generalizations or theories about these factors.
1. What are the key characteristics of research report?
You May Also Like:
Scope of Business Research
Data Collection
Questionnaire
Difference between questionnaire and schedule
Measurement
Data Processing
Nature of Research
Leave a Comment Cancel Reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.

- Research Report
- Post last modified: 11 January 2022
- Reading time: 25 mins read
- Post category: Research Methodology

What is Research Report?
Research reporting is the oral or written presentation of the findings in such detail and form as to be readily understood and assessed by the society, economy or particularly by the researchers.
As earlier said that it is the final stage of the research process and its purpose is to convey to interested persons the whole result of the study. Report writing is common to both academic and managerial situations. In academics, a research report is prepared for comprehensive and application-oriented learning. In businesses or organisations, reports are used for the basis of decision making.
Table of Content
- 1 What is Research Report?
- 2 Research Report Definition
- 3.1 Preliminary Part
- 3.2 Introduction of the Report
- 3.3 Review of Literature
- 3.4 The Research Methodology
- 3.5 Results
- 3.6 Concluding Remarks
- 3.7 Bibliography
- 4 Significance of Report Writing
- 5 Qualities of Good Report
- 6.1 Analysis of the subject matter
- 6.2 Research outline
- 6.3 Preparation of rough draft
- 6.4 Rewriting and polishing
- 6.5 Writing the final draft
- 7 Precautions for Writing Research Reports
- 8.1.1 Technical Report
- 8.1.2 Popular Report
- 8.2.1 Written Report
- 8.2.2 Oral Report
Research Report Definition
According to C. A. Brown , “A report is a communication from someone who has information to someone who wants to use that information.”
According to Goode and Hatt , “The preparation of report is the final stage of research, and it’s purpose is to convey to the interested persons the whole result of the study, in sufficient detail and so arranged as to enable each reader to comprehend the data and to determine for himself the validity of the conclusions.”
It is clear from the above definitions of a research report, it is a brief account of the problem of investigation, the justification of its selection and the procedure of analysis and interpretation. It is only a summary of the entire research proceedings.
In other words, it can be defined as written documents, which presents information in a specialized and concise manner.
Contents of Research Report
Although no hard and fast rules can be laid down, the report must contain the following points.
- Acknowledgement
- Table of contents
- List of tables
- List of graphs
- Introduction
- Background of the research study
- Statement of the problem
- Brief outline of the chapters
- Books review
- Review of articles published in books, journals, periodicals, etc
- Review of articles published in leading newspapers
- Working papers / discusssion paper / study reports
- Articles on authorised websites
- A broad conclusion and indications for further research
- The theoretical framework (variables)
- Model / hypothesis
- Instruments for data collection
- Data collection
- Pilot study
- Processing of data
- Hypothesis / model testing
- Data analysis and interpretation
- Tables and figures
- Conclusions
- Shortcomings
- Suggestions to the problems
- Direction for further research
Preliminary Part
The preliminary part may have seven major components – cover, title, preface, acknowledgement, table of contents, list of tables, list of graphs. Long reports presented in book form have a cover made up of a card sheet. The cover contains title of the research report, the authority to whom the report is submitted, name of the author, etc.
The preface introduces the report to the readers. It gives a very brief introduction of the report. In the acknowledgements author mention names of persons and organisations that have extended co-operation and helped in the various stages of research. Table of contents is essential. It gives the title and page number of each chapter.
Introduction of the Report
The introduction of the research report should clearly and logically bring out the background of the problem addressed in the research. The purpose of the introduction is to introduce the research project to the readers. A clear statement of the problem with specific questions to be answered is presented in the introduction. It contains a brief outline of the chapters.
Review of Literature
The third section reviews the important literature related to the study. A comprehensive review of the research literature referred to must be made. Previous research studies and the important writings in the area under study should be reviewed. Review of literature is helpful to provide a background for the development of the present study.
The researcher may review concerned books, articles published in edited books, journals and periodicals. Researcher may also take review of articles published in leading newspapers. A researcher should study working papers/discussion papers/study reports. It is essential for a broad conclusion and indications for further research.
The Research Methodology
Research methodology is an integral part of the research. It should clearly indicate the universe and the selection of samples, techniques of data collection, analysis and interpretation, statistical techniques, etc.
Results contain pilot study, processing of data, hypothesis/model testing, data analysis and interpretation, tables and figures, etc. This is the heart of the research report. If a pilot study is planned to be used, it’s purpose should be given in the research methodology.
The collected data and the information should be edited, coded, tabulated and analysed with a view to arriving at a valid and authentic conclusion. Tables and figures are used to clarify the significant relationship. The results obtained through tables, graphs should be critically interpreted.
Concluding Remarks
The concluding remarks should discuss the results obtained in the earlier sections, as well as their usefulness and implications. It contains findings, conclusions, shortcomings, suggestions to the problem and direction for future research. Findings are statements of factual information based upon the data analysis.
Conclusions must clearly explain whether the hypothesis have been established and rejected. This part requires great expertise and preciseness. A report should also refer to the limitations of the applicability of the research inferences. It is essential to suggest the theoretical, practical and policy implications of the research. The suggestions should be supported by scientific and logical arguments. The future direction of research based on the work completed should also be outlined.
Bibliography
The bibliography is an alphabetic list of books, journal articles, reports, etc, published or unpublished, read, referred to, examined by the researcher in preparing the report. The bibliography should follow standard formats for books, journal articles, research reports.
The end of the research report may consist of appendices, listed in respect of all technical data. Appendices are for the purpose of providing detailed data or information that would be too cumbersome within the main body of the research report.
Significance of Report Writing
Report writing is an important communication medium in organisations. The most crucial findings might have come out through a research report. Report is common to academics and managers also. Reports are used for comprehensive and application oriented learning in academics. In organisations, reports are used for the basis of decision making. The importance of report writing can be discussed as under.
Through research reports, a manager or an executive can quickly get an idea of a current scenario which improves his information base for making sound decisions affecting future operations of the company or enterprise. The research report acts as a means of communication of various research findings to the interested parties, organisations and general public.
Good report writing play, a significant role of conveying unknown facts about the phenomenon to the concerned parties. This may provide new insights and new opportunities to the people. Research report plays a key role in making effective decisions in marketing, production, banking, materials, human resource development and government also. Good report writing is used for economic planning and optimum utilisation of resources for the development of a nation.
Report writing facilitates the validation of generalisation. A research report is an end product of research. As earlier said that report writing provides useful information in arriving at rational decisions that may reform the business and society. The findings, conclusions, suggestions and recommendations are useful to academicians, scholars and policymakers. Report writing provides reference material for further research in the same or similar areas of research to the concerned parties.
While preparing a research report, a researcher should take some proper precautions. Report writing should be simple, lucid and systematic. Report writing should be written speedily without interrupting the continuity of thought. The report writing should sustain the interest of readers.
Qualities of Good Report
Report writing is a highly skilled job. It is a process of analysing, understanding and consolidating the findings and projecting a meaningful view of the phenomenon studied. A good report writing is essential for effective communication.
Following are the essential qualities of good report:
- A research report is essentially a scientific documentation. It should have a suggestive title, headings and sub-headings, paragraphs arranged in a logical sequence.
- Good research report should include everything that is relevant and exclude everything that is irrelevant. It means that it should contain the facts rather than opinion.
- The language of the report should be simple and unambiguous. It means that it should be free from biases of the researchers derived from the past experience. Confusion, pretentiousness and pomposity should be carefully guarded against. It means that the language of the report should be simple, employing appropriate words, idioms and expressions.
- The report must be free from grammatical mistakes. It must be grammatically accurate. Faulty construction of sentences makes the meaning of the narrative obscure and ambiguous.
- The report has to take into consideration two facts. Firstly, for whom the report is meant and secondly, what is his level of knowledge. The report has to look to the subject matter of the report and the fact as to the level of knowledge of the person for whom it is meant. Because all reports are not meant for research scholars.
Steps in Writing Research Report
Report writing is a time consuming and expensive exercise. Therefore, reports have to be very sharply focused in purpose content and readership. There is no single universally acceptable method of writing a research report.
Following are the general steps in writing a research report:
Analysis of the subject matter
Research outline, preparation of rough draft, rewriting and polishing, writing the final draft.
This is the first and important step in writing a research report. It is concerned with the development of a subject. Subject matter should be written in a clear, logical and concise manner. The style adopted should be open, straightforward and dignified and folk style language should be avoided.
The data, the reliability and validity of the results of the statistical analysis should be in the form of tables, figures and equations. All redundancy in the data or results presented should be eliminated.
The research outline is an organisational framework prepared by the researcher well in advance. It is an aid to logical organisation of material and a reminder of the points to be stressed in the report. In the process of writing, if need be, outline may be revised accordingly.
Time and place of the study, scope and limitations of the study, study design, summary of pilot study, methods of data collection, analysis interpretation, etc., may be included in a research outline.
Having prepared the primary and secondary data, the researcher has to prepare a rough draft. While preparing the rough draft, the researcher should keep the objectives of the research in mind, and focus on one objective at a time. The researcher should make a checklist of the important points that are necessary to be covered in the manuscript. A researcher should use dictionary and relevant reference materials as and when required.
This is an important step in writing a research report. It takes more time than a rough draft. While rewriting and polishing, a researcher should check the report for weakness in logical development or presentation. He should take breaks in between rewriting and polishing since this gives the time to incubate the ideas.
The last and important step is writing the final draft. The language of the report should be simple, employing appropriate words and expressions and should avoid vague expressions such as ‘it seems’ and ‘there may be’ etc.
It should not used personal pronouns, such as I, We, My, Us, etc and should substitute these by such expressions as a researcher, investigator, etc. Before the final drafting of the report, it is advisable that the researcher should prepare a first draft for critical considerations and possible improvements. It will be helpful in writing the final draft. Finally, the report should be logically outlined with the future directions of the research based on the work completed.
Precautions for Writing Research Reports
A research report is a means of conveying the research study to a specific target audience. The following precautions should be taken while preparing a research report:
- Its hould belong enough to cover the subject and short enough to preserve interest.
- It should not be dull and complicated.
- It should be simple, without the usage of abstract terms and technical jargons.
- It should offer ready availability of findings with the help of charts, tables and graphs, as readers prefer quick knowledge of main findings.
- The layout of the report should be in accordance with the objectives of the research study.
- There should be no grammatical errors and writing should adhere to the techniques of report writing in case of quotations, footnotes and documentations.
- It should be original, intellectual and contribute to the solution of a problem or add knowledge to the concerned field.
- Appendices should been listed with respect to all the technical data in the report.
- It should be attractive, neat and clean, whether handwritten or typed.
- The report writer should refrain from confusing the possessive form of the word ‘it’ is with ‘it’s.’ The accurate possessive form of ‘it is’ is ‘its.’ The use of ‘it’s’ is the contractive form of ‘it is.
- A report should not have contractions. Examples are ‘didn’t’ or ‘it’s.’ In report writing, it is best to use the non-contractive form. Therefore, the examples would be replaced by ‘did not’ and ‘it is.’ Using ‘Figure’ instead of ‘Fig.’ and ‘Table’ instead of ‘Tab.’ will spare the reader of having to translate the abbreviations, while reading. If abbreviations are used, use them consistently throughout the report. For example, do not switch among ‘versus,’ and ‘vs’.
- It is advisable to avoid using the word ‘very’ and other such words that try to embellish a description. They do not add any extra meaning and, therefore, should be dropped.
- Repetition hampers lucidity. Report writers must avoid repeating the same word more than once within a sentence.
- When you use the word ‘this’ or ‘these’ make sure you indicate to what you are referring. This reduces the ambiguity in your writing and helps to tie sentences together.
- Do not use the word ‘they’ to refer to a singular person. You can either rewrite the sentence to avoid needing such a reference or use the singular ‘he or she.’
Types of Research Report
Research reports are designed in order to convey and record the information that will be of practical use to the reader. It is organized into distinct units of specific and highly visible information. The kind of audience addressed in the research report decides the type of report.
Research reports can be categorized on the following basis:
Classification on the Basis of Information
Classification on the basis of representation.
Following are the ways through which the results of the research report can be presented on the basis of information contained:
Technical Report
A technical report is written for other researchers. In writing the technical reports, the importance is mainly given to the methods that have been used to collect the information and data, the presumptions that are made and finally, the various presentation techniques that are used to present the findings and data.
Following are main features of a technical report:
- Summary: It covers a brief analysis of the findings of the research in a very few pages.
- Nature: It contains the reasons for which the research is undertaken, the analysis and the data that is required in order to prepare a report.
- Methods employed: It contains a description of the methods that were employed in order to collect the data.
- Data: It covers a brief analysis of the various sources from which the data has been collected with their features and drawbacks
- Analysis of data and presentation of the findings: It contains the various forms through which the data that has been analysed can be presented.
- Conclusions: It contains a brief explanation of findings of the research.
- Bibliography: It contains a detailed analysis of the various bibliographies that have been used in order to conduct a research.
- Technical appendices: It contains the appendices for the technical matters and for questionnaires and mathematical derivations.
- Index: The index of the technical report must be provided at the end of the report.
Popular Report
A popular report is formulated when there is a need to draw conclusions of the findings of the research report. One of the main points of consideration that should be kept in mind while formulating a research report is that it must be simple and attractive. It must be written in a very simple manner that is understandable to all. It must also be made attractive by using large prints, various sub-headings and by giving cartoons occasionally.
Following are the main points that must be kept in mind while preparing a popular report:
- Findings and their implications : While preparing a popular report, main importance is given to the findings of the information and the conclusions that can be drawn out of these findings.
- Recommendations for action : If there are any deviations in the report then recommendations are made for taking corrective action in order to rectify the errors.
- Objective of the study : In a popular report, the specific objective for which the research has been undertaken is presented.
- Methods employed : The report must contain the various methods that has been employed in order to conduct a research.
- Results : The results of the research findings must be presented in a suitable and appropriate manner by taking the help of charts and diagrams.
- Technical appendices : The report must contain an in-depth information used to collect the data in the form of appendices.
Following are the ways through which the results of the research report can be presented on the basis of representation:
- Writtenreport
- Oral report
Written Report
A written report plays a vital role in every business operation. The manner in which an organization writes business letters and business reports creates an impression of its standard. Therefore, the organization should emphasize on the improvement of the writing skills of the employees in order to maintain effective relations with their customers.
Writing effective written reports requires a lot of hard work. Therefore, before you begin writing, it is important to know the objective, i.e., the purpose of writing, collection and organization of required data.
Oral Report
At times, oral presentation of the results that are drawn out of research is considered effective, particularly in cases where policy recommendations are to be made. This approach proves beneficial because it provides a medium of interaction between a listener and a speaker. This leads to a better understanding of the findings and their implications.
However, the main drawback of oral presentation is the lack of any permanent records related to the research. Oral presentation of the report is also effective when it is supported with various visual devices, such as slides, wall charts and whiteboards that help in better understanding of the research reports.
Business Ethics
( Click on Topic to Read )
- What is Ethics?
- What is Business Ethics?
- Values, Norms, Beliefs and Standards in Business Ethics
- Indian Ethos in Management
- Ethical Issues in Marketing
- Ethical Issues in HRM
- Ethical Issues in IT
- Ethical Issues in Production and Operations Management
- Ethical Issues in Finance and Accounting
- What is Corporate Governance?
- What is Ownership Concentration?
- What is Ownership Composition?
- Types of Companies in India
- Internal Corporate Governance
- External Corporate Governance
- Corporate Governance in India
- What is Enterprise Risk Management (ERM)?
- What is Assessment of Risk?
- What is Risk Register?
- Risk Management Committee
Corporate social responsibility (CSR)
- Theories of CSR
- Arguments Against CSR
- Business Case for CSR
- Importance of CSR in India
- Drivers of Corporate Social Responsibility
- Developing a CSR Strategy
- Implement CSR Commitments
- CSR Marketplace
- CSR at Workplace
- Environmental CSR
- CSR with Communities and in Supply Chain
- Community Interventions
- CSR Monitoring
- CSR Reporting
- Voluntary Codes in CSR
- What is Corporate Ethics?
Lean Six Sigma
- What is Six Sigma?
- What is Lean Six Sigma?
- Value and Waste in Lean Six Sigma
- Six Sigma Team
- MAIC Six Sigma
- Six Sigma in Supply Chains
- What is Binomial, Poisson, Normal Distribution?
- What is Sigma Level?
- What is DMAIC in Six Sigma?
- What is DMADV in Six Sigma?
- Six Sigma Project Charter
- Project Decomposition in Six Sigma
- Critical to Quality (CTQ) Six Sigma
- Process Mapping Six Sigma
- Flowchart and SIPOC
- Gage Repeatability and Reproducibility
- Statistical Diagram
- Lean Techniques for Optimisation Flow
- Failure Modes and Effects Analysis (FMEA)
- What is Process Audits?
- Six Sigma Implementation at Ford
- IBM Uses Six Sigma to Drive Behaviour Change
- Research Methodology
- What is Research?
What is Hypothesis?
- Sampling Method
Research Methods
- Data Collection in Research
Methods of Collecting Data
- Application of Business Research
Levels of Measurement
- What is Sampling?
- Hypothesis Testing
- What is Management?
- Planning in Management
- Decision Making in Management
- What is Controlling?
- What is Coordination?
- What is Staffing?
- Organization Structure
- What is Departmentation?
- Span of Control
- What is Authority?
- Centralization vs Decentralization
- Organizing in Management
- Schools of Management Thought
- Classical Management Approach
- Is Management an Art or Science?
- Who is a Manager?
Operations Research
- What is Operations Research?
- Operation Research Models
- Linear Programming
- Linear Programming Graphic Solution
- Linear Programming Simplex Method
- Linear Programming Artificial Variable Technique
- Duality in Linear Programming
- Transportation Problem Initial Basic Feasible Solution
- Transportation Problem Finding Optimal Solution
- Project Network Analysis with Critical Path Method
- Project Network Analysis Methods
- Project Evaluation and Review Technique (PERT)
- Simulation in Operation Research
- Replacement Models in Operation Research
Operation Management
- What is Strategy?
- What is Operations Strategy?
- Operations Competitive Dimensions
- Operations Strategy Formulation Process
- What is Strategic Fit?
- Strategic Design Process
- Focused Operations Strategy
- Corporate Level Strategy
- Expansion Strategies
- Stability Strategies
- Retrenchment Strategies
- Competitive Advantage
- Strategic Choice and Strategic Alternatives
- What is Production Process?
- What is Process Technology?
- What is Process Improvement?
- Strategic Capacity Management
- Production and Logistics Strategy
- Taxonomy of Supply Chain Strategies
- Factors Considered in Supply Chain Planning
- Operational and Strategic Issues in Global Logistics
- Logistics Outsourcing Strategy
- What is Supply Chain Mapping?
- Supply Chain Process Restructuring
- Points of Differentiation
- Re-engineering Improvement in SCM
- What is Supply Chain Drivers?
- Supply Chain Operations Reference (SCOR) Model
- Customer Service and Cost Trade Off
- Internal and External Performance Measures
- Linking Supply Chain and Business Performance
- Netflix’s Niche Focused Strategy
- Disney and Pixar Merger
- Process Planning at Mcdonald’s
Service Operations Management
- What is Service?
- What is Service Operations Management?
- What is Service Design?
- Service Design Process
- Service Delivery
- What is Service Quality?
- Gap Model of Service Quality
- Juran Trilogy
- Service Performance Measurement
- Service Decoupling
- IT Service Operation
- Service Operations Management in Different Sector
Procurement Management
- What is Procurement Management?
- Procurement Negotiation
- Types of Requisition
- RFX in Procurement
- What is Purchasing Cycle?
- Vendor Managed Inventory
- Internal Conflict During Purchasing Operation
- Spend Analysis in Procurement
- Sourcing in Procurement
- Supplier Evaluation and Selection in Procurement
- Blacklisting of Suppliers in Procurement
- Total Cost of Ownership in Procurement
- Incoterms in Procurement
- Documents Used in International Procurement
- Transportation and Logistics Strategy
- What is Capital Equipment?
- Procurement Process of Capital Equipment
- Acquisition of Technology in Procurement
- What is E-Procurement?
- E-marketplace and Online Catalogues
- Fixed Price and Cost Reimbursement Contracts
- Contract Cancellation in Procurement
- Ethics in Procurement
- Legal Aspects of Procurement
- Global Sourcing in Procurement
- Intermediaries and Countertrade in Procurement
Strategic Management
- What is Strategic Management?
- What is Value Chain Analysis?
- Mission Statement
- Business Level Strategy
- What is SWOT Analysis?
- What is Competitive Advantage?
- What is Vision?
- What is Ansoff Matrix?
- Prahalad and Gary Hammel
- Strategic Management In Global Environment
- Competitor Analysis Framework
- Competitive Rivalry Analysis
- Competitive Dynamics
- What is Competitive Rivalry?
- Five Competitive Forces That Shape Strategy
- What is PESTLE Analysis?
- Fragmentation and Consolidation Of Industries
- What is Technology Life Cycle?
- What is Diversification Strategy?
- What is Corporate Restructuring Strategy?
- Resources and Capabilities of Organization
- Role of Leaders In Functional-Level Strategic Management
- Functional Structure In Functional Level Strategy Formulation
- Information And Control System
- What is Strategy Gap Analysis?
- Issues In Strategy Implementation
- Matrix Organizational Structure
- What is Strategic Management Process?
Supply Chain
- What is Supply Chain Management?
- Supply Chain Planning and Measuring Strategy Performance
- What is Warehousing?
- What is Packaging?
- What is Inventory Management?
- What is Material Handling?
- What is Order Picking?
- Receiving and Dispatch, Processes
- What is Warehouse Design?
- What is Warehousing Costs?
You Might Also Like
What is sample size determination, formula, determining,, what is causal research advantages, disadvantages, how to perform, what is hypothesis testing procedure, what is hypothesis definition, meaning, characteristics, sources, what is measure of skewness, types of charts used in data analysis, primary data and secondary data, what is research types, purpose, characteristics, process, leave a reply cancel reply.
You must be logged in to post a comment.
World's Best Online Courses at One Place
We’ve spent the time in finding, so you can spend your time in learning
Digital Marketing
Personal Growth

Development
Research Methodology WRITING A RESEARCH REPORT
- September 2021
- In book: Research Methodology in Social Sciences (A Short Manual) (pp.177)
- Publisher: New Delhi: Corvette Press

- Himachal Pradesh University
Discover the world's research
- 25+ million members
- 160+ million publication pages
- 2.3+ billion citations
- Miriam Hill
- Derek Jansen
- Kerryn Warren
- Kretchmer Paul
- Recruit researchers
- Join for free
- Login Email Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google Welcome back! Please log in. Email · Hint Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google No account? Sign up
- Privacy Policy

Home » Research Results Section – Writing Guide and Examples
Research Results Section – Writing Guide and Examples
Table of Contents

Research Results
Research results refer to the findings and conclusions derived from a systematic investigation or study conducted to answer a specific question or hypothesis. These results are typically presented in a written report or paper and can include various forms of data such as numerical data, qualitative data, statistics, charts, graphs, and visual aids.
Results Section in Research
The results section of the research paper presents the findings of the study. It is the part of the paper where the researcher reports the data collected during the study and analyzes it to draw conclusions.
In the results section, the researcher should describe the data that was collected, the statistical analysis performed, and the findings of the study. It is important to be objective and not interpret the data in this section. Instead, the researcher should report the data as accurately and objectively as possible.
Structure of Research Results Section
The structure of the research results section can vary depending on the type of research conducted, but in general, it should contain the following components:
- Introduction: The introduction should provide an overview of the study, its aims, and its research questions. It should also briefly explain the methodology used to conduct the study.
- Data presentation : This section presents the data collected during the study. It may include tables, graphs, or other visual aids to help readers better understand the data. The data presented should be organized in a logical and coherent way, with headings and subheadings used to help guide the reader.
- Data analysis: In this section, the data presented in the previous section are analyzed and interpreted. The statistical tests used to analyze the data should be clearly explained, and the results of the tests should be presented in a way that is easy to understand.
- Discussion of results : This section should provide an interpretation of the results of the study, including a discussion of any unexpected findings. The discussion should also address the study’s research questions and explain how the results contribute to the field of study.
- Limitations: This section should acknowledge any limitations of the study, such as sample size, data collection methods, or other factors that may have influenced the results.
- Conclusions: The conclusions should summarize the main findings of the study and provide a final interpretation of the results. The conclusions should also address the study’s research questions and explain how the results contribute to the field of study.
- Recommendations : This section may provide recommendations for future research based on the study’s findings. It may also suggest practical applications for the study’s results in real-world settings.
Outline of Research Results Section
The following is an outline of the key components typically included in the Results section:
I. Introduction
- A brief overview of the research objectives and hypotheses
- A statement of the research question
II. Descriptive statistics
- Summary statistics (e.g., mean, standard deviation) for each variable analyzed
- Frequencies and percentages for categorical variables
III. Inferential statistics
- Results of statistical analyses, including tests of hypotheses
- Tables or figures to display statistical results
IV. Effect sizes and confidence intervals
- Effect sizes (e.g., Cohen’s d, odds ratio) to quantify the strength of the relationship between variables
- Confidence intervals to estimate the range of plausible values for the effect size
V. Subgroup analyses
- Results of analyses that examined differences between subgroups (e.g., by gender, age, treatment group)
VI. Limitations and assumptions
- Discussion of any limitations of the study and potential sources of bias
- Assumptions made in the statistical analyses
VII. Conclusions
- A summary of the key findings and their implications
- A statement of whether the hypotheses were supported or not
- Suggestions for future research
Example of Research Results Section
An Example of a Research Results Section could be:
- This study sought to examine the relationship between sleep quality and academic performance in college students.
- Hypothesis : College students who report better sleep quality will have higher GPAs than those who report poor sleep quality.
- Methodology : Participants completed a survey about their sleep habits and academic performance.
II. Participants
- Participants were college students (N=200) from a mid-sized public university in the United States.
- The sample was evenly split by gender (50% female, 50% male) and predominantly white (85%).
- Participants were recruited through flyers and online advertisements.
III. Results
- Participants who reported better sleep quality had significantly higher GPAs (M=3.5, SD=0.5) than those who reported poor sleep quality (M=2.9, SD=0.6).
- See Table 1 for a summary of the results.
- Participants who reported consistent sleep schedules had higher GPAs than those with irregular sleep schedules.
IV. Discussion
- The results support the hypothesis that better sleep quality is associated with higher academic performance in college students.
- These findings have implications for college students, as prioritizing sleep could lead to better academic outcomes.
- Limitations of the study include self-reported data and the lack of control for other variables that could impact academic performance.
V. Conclusion
- College students who prioritize sleep may see a positive impact on their academic performance.
- These findings highlight the importance of sleep in academic success.
- Future research could explore interventions to improve sleep quality in college students.
Example of Research Results in Research Paper :
Our study aimed to compare the performance of three different machine learning algorithms (Random Forest, Support Vector Machine, and Neural Network) in predicting customer churn in a telecommunications company. We collected a dataset of 10,000 customer records, with 20 predictor variables and a binary churn outcome variable.
Our analysis revealed that all three algorithms performed well in predicting customer churn, with an overall accuracy of 85%. However, the Random Forest algorithm showed the highest accuracy (88%), followed by the Support Vector Machine (86%) and the Neural Network (84%).
Furthermore, we found that the most important predictor variables for customer churn were monthly charges, contract type, and tenure. Random Forest identified monthly charges as the most important variable, while Support Vector Machine and Neural Network identified contract type as the most important.
Overall, our results suggest that machine learning algorithms can be effective in predicting customer churn in a telecommunications company, and that Random Forest is the most accurate algorithm for this task.
Example 3 :
Title : The Impact of Social Media on Body Image and Self-Esteem
Abstract : This study aimed to investigate the relationship between social media use, body image, and self-esteem among young adults. A total of 200 participants were recruited from a university and completed self-report measures of social media use, body image satisfaction, and self-esteem.
Results: The results showed that social media use was significantly associated with body image dissatisfaction and lower self-esteem. Specifically, participants who reported spending more time on social media platforms had lower levels of body image satisfaction and self-esteem compared to those who reported less social media use. Moreover, the study found that comparing oneself to others on social media was a significant predictor of body image dissatisfaction and lower self-esteem.
Conclusion : These results suggest that social media use can have negative effects on body image satisfaction and self-esteem among young adults. It is important for individuals to be mindful of their social media use and to recognize the potential negative impact it can have on their mental health. Furthermore, interventions aimed at promoting positive body image and self-esteem should take into account the role of social media in shaping these attitudes and behaviors.
Importance of Research Results
Research results are important for several reasons, including:
- Advancing knowledge: Research results can contribute to the advancement of knowledge in a particular field, whether it be in science, technology, medicine, social sciences, or humanities.
- Developing theories: Research results can help to develop or modify existing theories and create new ones.
- Improving practices: Research results can inform and improve practices in various fields, such as education, healthcare, business, and public policy.
- Identifying problems and solutions: Research results can identify problems and provide solutions to complex issues in society, including issues related to health, environment, social justice, and economics.
- Validating claims : Research results can validate or refute claims made by individuals or groups in society, such as politicians, corporations, or activists.
- Providing evidence: Research results can provide evidence to support decision-making, policy-making, and resource allocation in various fields.
How to Write Results in A Research Paper
Here are some general guidelines on how to write results in a research paper:
- Organize the results section: Start by organizing the results section in a logical and coherent manner. Divide the section into subsections if necessary, based on the research questions or hypotheses.
- Present the findings: Present the findings in a clear and concise manner. Use tables, graphs, and figures to illustrate the data and make the presentation more engaging.
- Describe the data: Describe the data in detail, including the sample size, response rate, and any missing data. Provide relevant descriptive statistics such as means, standard deviations, and ranges.
- Interpret the findings: Interpret the findings in light of the research questions or hypotheses. Discuss the implications of the findings and the extent to which they support or contradict existing theories or previous research.
- Discuss the limitations : Discuss the limitations of the study, including any potential sources of bias or confounding factors that may have affected the results.
- Compare the results : Compare the results with those of previous studies or theoretical predictions. Discuss any similarities, differences, or inconsistencies.
- Avoid redundancy: Avoid repeating information that has already been presented in the introduction or methods sections. Instead, focus on presenting new and relevant information.
- Be objective: Be objective in presenting the results, avoiding any personal biases or interpretations.

When to Write Research Results
Here are situations When to Write Research Results”
- After conducting research on the chosen topic and obtaining relevant data, organize the findings in a structured format that accurately represents the information gathered.
- Once the data has been analyzed and interpreted, and conclusions have been drawn, begin the writing process.
- Before starting to write, ensure that the research results adhere to the guidelines and requirements of the intended audience, such as a scientific journal or academic conference.
- Begin by writing an abstract that briefly summarizes the research question, methodology, findings, and conclusions.
- Follow the abstract with an introduction that provides context for the research, explains its significance, and outlines the research question and objectives.
- The next section should be a literature review that provides an overview of existing research on the topic and highlights the gaps in knowledge that the current research seeks to address.
- The methodology section should provide a detailed explanation of the research design, including the sample size, data collection methods, and analytical techniques used.
- Present the research results in a clear and concise manner, using graphs, tables, and figures to illustrate the findings.
- Discuss the implications of the research results, including how they contribute to the existing body of knowledge on the topic and what further research is needed.
- Conclude the paper by summarizing the main findings, reiterating the significance of the research, and offering suggestions for future research.
Purpose of Research Results
The purposes of Research Results are as follows:
- Informing policy and practice: Research results can provide evidence-based information to inform policy decisions, such as in the fields of healthcare, education, and environmental regulation. They can also inform best practices in fields such as business, engineering, and social work.
- Addressing societal problems : Research results can be used to help address societal problems, such as reducing poverty, improving public health, and promoting social justice.
- Generating economic benefits : Research results can lead to the development of new products, services, and technologies that can create economic value and improve quality of life.
- Supporting academic and professional development : Research results can be used to support academic and professional development by providing opportunities for students, researchers, and practitioners to learn about new findings and methodologies in their field.
- Enhancing public understanding: Research results can help to educate the public about important issues and promote scientific literacy, leading to more informed decision-making and better public policy.
- Evaluating interventions: Research results can be used to evaluate the effectiveness of interventions, such as treatments, educational programs, and social policies. This can help to identify areas where improvements are needed and guide future interventions.
- Contributing to scientific progress: Research results can contribute to the advancement of science by providing new insights and discoveries that can lead to new theories, methods, and techniques.
- Informing decision-making : Research results can provide decision-makers with the information they need to make informed decisions. This can include decision-making at the individual, organizational, or governmental levels.
- Fostering collaboration : Research results can facilitate collaboration between researchers and practitioners, leading to new partnerships, interdisciplinary approaches, and innovative solutions to complex problems.
Advantages of Research Results
Some Advantages of Research Results are as follows:
- Improved decision-making: Research results can help inform decision-making in various fields, including medicine, business, and government. For example, research on the effectiveness of different treatments for a particular disease can help doctors make informed decisions about the best course of treatment for their patients.
- Innovation : Research results can lead to the development of new technologies, products, and services. For example, research on renewable energy sources can lead to the development of new and more efficient ways to harness renewable energy.
- Economic benefits: Research results can stimulate economic growth by providing new opportunities for businesses and entrepreneurs. For example, research on new materials or manufacturing techniques can lead to the development of new products and processes that can create new jobs and boost economic activity.
- Improved quality of life: Research results can contribute to improving the quality of life for individuals and society as a whole. For example, research on the causes of a particular disease can lead to the development of new treatments and cures, improving the health and well-being of millions of people.
About the author
Muhammad Hassan
Researcher, Academic Writer, Web developer
You may also like

Research Report – Example, Writing Guide and...

References in Research – Types, Examples and...

Thesis Outline – Example, Template and Writing...

Ethical Considerations – Types, Examples and...

Research Objectives – Types, Examples and...

APA Research Paper Format – Example, Sample and...

Types of Research Reports

What is a Research Report?
A research report is a concise document that summarizes the findings, methods, and conclusions of a research study or investigation. There are various types of research reports available for different purposes.
It typically includes details on the research question, methodology, data analysis, and results, providing a structured and informative account of the research process and outcomes.

Table of Contents
- Market Research
- Experimental
- Descriptive
- Exploratory
- Explanatory
Types of Research Report Writing
Limitations, key highlights.
- A research report is a document that gives a quick overview of a research study.
- Types of research reports offer a standardized format and structure, making it easier for readers to navigate and comprehend the information.
- They are useful in fields like academia, business, healthcare, social sciences, and more.
- Different types of report writing determine the report’s primary purpose, i.e., if it should be short, long, or for internal purposes, etc.
Different Types Of Research Reports
1. technical or scientific reports.
Technical and scientific reports communicate research findings to experts and professionals in a particular field.
Start Your Free Data Science Course
Hadoop, Data Science, Statistics & others
Characteristics:
- These reports include technical jargon, detailed methodologies, and in-depth analysis.
- They often have a standardized format for peer review.
2. Popular Reports
Popular reports are designed for a general audience and aim to inform, educate, or entertain on a wide range of topics.
- Includes clear and jargon-free language
- Uses storytelling, visuals, and anecdotes to engage readers
- Prioritizes readability over deep analysis.
3. Survey Reports
Survey reports include data collected through surveys and focus on presenting insights and opinions on specific issues or questions.
- Contains information on survey methodology, including sample size and data collection methods.
- Presents statistical summaries like percentages and charts.
4. Market Research Reports
Market research reports provide insights into consumer behavior, market trends, and industry analysis.
- Includes market surveys, competitor analysis, and consumer demographics.
- Helps businesses in making strategic decisions.
5. Case Study Reports
Case study reports focus on an in-depth examination of a single entity, often to explore complex, real-life situations.
- Includes detailed descriptions of the case, data collection methods, and analysis.
- Common in business and psychology fields.
6. Analytical Research Reports
Analytical research reports involve a deep analysis of data to uncover patterns, trends, or relationships.
- Uses statistical tools and software to analyze data comprehensively
- Common in fields like economics and social sciences.
7. Review or Literature Survey Reports
Literature review reports provide an overview of existing research on a specific topic, highlighting gaps and trends.
- Synthesizes findings from various sources and provides a historical context
- Often offers recommendations for future research.
8. Experimental Research Reports
Experimental research reports involve controlled experiments to test hypotheses and determine if the results support or reject the hypothesis.
- Uses random sampling and control groups to minimize bias.
- Includes detailed descriptions of the experiment, hypothesis, methods, and statistical analyses.
9. Descriptive Research Reports
Descriptive research reports aim to provide a comprehensive picture of a phenomenon, group, or situation. They seek to answer the “what” and “how” questions.
- Typically, it relies on observations, surveys, and content analysis.
- Focuses on describing and summarizing data.
10. Exploratory Research Reports
Exploratory research reports are conducted when there is little prior knowledge about a subject. They aim to identify key variables and research questions.
- Involves open-ended interviews, focus groups, and literature reviews.
- Findings are preliminary, serving as a basis for further research.
11. Explanatory Research Reports
Explanatory research reports seek to understand the relationships between variables and explain why certain phenomena occur.
- Uses experimental designs, surveys, and statistical analyses.
- Provides in-depth insights into the research problem.
12. Policy or White Papers
Policy or white papers aim to inform policymakers, stakeholders, and the public about specific issues and recommend actions.
- Presents research findings in a concise and accessible manner
- Often consists of policy recommendations.
Components of Research Reports
These are some common components you must know while writing different types of research reports.
1. Title Page:
- Title of the Report
- Institutional Affiliation
2. Abstract: Add a concise summary of the research, including the research question or objective, methodology, key findings, and implications. Typically, it should be no more than 150-250 words.
3. Table of Contents: Include a list of sections and subsections with page numbers.
4. List of Figures and Tables: If your research includes numerical data, add all the statistics and tables along with their corresponding page numbers. It is similar to a table of contents for quantitative data.
5. List of Abbreviations and Symbols: Include any abbreviations or symbols you have used in the report and their meanings.
6. Introduction:
- Provide background information on the topic.
- State the research question or objective.
- Explain the significance and purpose of the study.
- Provide an outline of the report’s structure.
7. Literature Review:
- Review relevant literature and previous research on the topic.
- Identify gaps in existing knowledge.
- Explain how your study contributes to the field.
8. Methodology:
- Describe the research methods and techniques that you used.
- Explain the sampling methods, data collection, and data analysis procedures.
- Discuss any ethical considerations.
9. Results:
- Present the findings of your research.
- Use tables, figures, charts, and graphs to illustrate key points.
- Include descriptive and inferential statistics as needed.
10. Discussion:
- Interpret the results and relate them to the research question.
- Discuss the implications of your findings.
- Compare your results to previous research.
- Address any limitations of your study.
11. Conclusion:
- Summarize the main findings and their significance.
- Restate the research question and how it was addressed.
- Suggest areas for future research.
12. References: Include a list of all the sources cited in your report in a standardized citation style (e.g., APA, MLA, Chicago).
Let us see an example of a research report.
Research Report: The Impact of Artificial Intelligence on the Labor Market
This research study explores the profound changes occurring in the labor market due to the increasing adoption of artificial intelligence (AI) technologies. The study examines the potential benefits and challenges AI poses for the workforce, job displacement, and the skills required in the future job market.
List of Figures and Tables
Introduction, literature review, methodology.
- Figure 1: Trend in AI Adoption by Industry (Page 7)
- Table 1: Summary of Job Displacement Data (Page 9)
- Figure 2: Projected Growth of AI-Related Occupations (Page 11)
The introduction section provides an overview of the research topic. It explains the significance of studying the impact of AI on the labor market, outlines the research questions, and previews the structure of the report.
The literature review section reviews existing research on the effects of AI on employment and the labor market. It discusses the different perspectives on whether AI will create new jobs or lead to job displacement. It also explores the skills and education required for the future workforce.
This section explains the research methods used, such as data collection methods, sources, and analytical techniques. It outlines how data on AI adoption, job displacement, and future job projections were gathered and analyzed.
The results section presents the key findings of the study. It includes data on the extent of AI adoption across industries, job displacement rates, and projections for AI-related occupations.
The discussion section interprets the results in the context of the research questions. It analyzes the potential benefits and challenges AI poses for the labor market, discusses policy implications, and explores the role of education and training in preparing the workforce for the AI era.
In conclusion, this research highlights the transformative impact of artificial intelligence on the labor market. While AI brings opportunities for innovation and efficiency, it also presents challenges related to job displacement and workforce adaptation. Preparing for this evolving job landscape is crucial for individuals and policymakers.
Given below are various types of research reports writing that researchers and organizations use to present findings, progress, and other information.
| Outlines a plan for a project or research for approval or funding. | Research proposal submitted to study the impact of climate change on local ecosystems. | |
| Generated at regular intervals to provide project updates. | Weekly sales reports summarizing product sales figures. | |
| Detailed, structured reports often used in academic, scientific, or business settings. | Formal business report analyzing a company’s financial performance for the year. | |
| Less structured reports for quick internal communication. | Email summarizing key takeaways from a team meeting. | |
| Concise documents offering a brief overview of a specific topic. | A one-page summary of customer feedback from a product launch. | |
| Comprehensive reports with in-depth analysis and information. | 100-page research report on the effects of a new drug on a medical condition. | |
| Focus on data analysis and provide insights or recommendations. | Market research report analyzing consumer behavior trends and recommending marketing strategies. | |
| Convey information without providing analysis or recommendations. | Report detailing the steps of a manufacturing process for new employees. | |
| Flow within the organizational hierarchy, moving up or down. | Report from a department manager to the company’s vice president on department performance. | |
| Sent between individuals or departments at the same organizational level. | Report from one project manager to another project manager in a different department. | |
| Created and distributed within an organization for internal purposes. | Internal audit report examining the company’s financial records for compliance. | |
| Prepared for external audiences, such as clients, investors, or regulators. | A publicly traded company publishes an annual report for shareholders and the general public. |
Here is why the different types of research reports are important.
- Research reports are a primary means of sharing new knowledge and insights with the academic and scientific community. They contribute to the growth of human understanding in various fields.
- They provide a detailed and structured account of the research process, including methods, data, analysis, and conclusions. This documentation is crucial for transparency, replication, and future reference.
- These reports hold researchers accountable for their work. They provide a transparent record of the study, allowing others to assess the validity and reliability of the research.
- These often influence policy decisions, business strategies, and practical applications. For instance, medical research informs healthcare practices, while market research guides business decisions.
Listed below are some limitations of different types of research reports.
- Research reports can be influenced by the researcher’s biases, preferences, or the funder’s interests. It’s essential to assess the methodology critically.
- Findings in research reports may not always be directly applicable to other contexts or populations.
- Certain research reports are not available to everyone due to several barriers, making it hard for people to access important information.
- The process of conducting research, writing a report, and getting it published can be time-consuming.
Final Thoughts
Different types of research reports are important for sharing knowledge, making smart choices, and moving forward in different areas of study. It’s vital for both researchers and those who use research to grasp the different kinds of reports, what goes into them, and why they matter.
Frequently Asked Questions (FAQs)
Q1. Are research reports the same as research papers? Answer: Research reports and research papers share similarities but have distinct purposes and structures. Research papers are often more academic and can vary in structure, while research reports are typically more structured and cater to a broader audience.
Q2. How do I choose the right type of research report for my study? Answer: The choice of research report type depends on your research goals, audience, and the nature of your study. Consider whether you are conducting scientific research, market analysis, academic research, or policy analysis, and select the format that aligns with your objectives.
Q3. Can research reports be used as references in other research reports? Answer: Yes, research reports can be cited and used as references in other research reports as long as they are credible sources. Citing previous research reports adds depth and credibility to your work.
Recommended Articles
This article lists all the types of research reports available for research methodologies. We have also included its format, example, and several report-writing methods. For similar articles, you can check the following articles,
- Types of Research Methodology
- Types of Quantitative Research
- Quantitative Research Examples
- What is Qualitative Data Analysis

*Please provide your correct email id. Login details for this Free course will be emailed to you
By signing up, you agree to our Terms of Use and Privacy Policy .
Forgot Password?
This website or its third-party tools use cookies, which are necessary to its functioning and required to achieve the purposes illustrated in the cookie policy. By closing this banner, scrolling this page, clicking a link or continuing to browse otherwise, you agree to our Privacy Policy

Explore 1000+ varieties of Mock tests View more
Submit Next Question
🚀 Limited Time Offer! - 🎁 ENROLL NOW

How to Write a Report: A Guide to Report Format and Best Practice
A report is a nonfiction account that presents and/or summarizes the facts about a particular event, topic, or issue. The idea is that people who are unfamiliar with the subject can find everything they need to know from a good report.
Reports make it easy to catch someone up to speed on a subject, but actually writing a report is anything but easy. So to help you understand what to do, below we present a little report of our own, all about report writing and report format best practices.
Communicate with confidence Grammarly helps you write the way you intend Write with Grammarly
Table of contents
What is a report?
Types of report formats
What is the structure of a report, what should be included in a report, how to write a report in 7 steps, what is a report .
In technical terms, the definition of a report is pretty vague: any account, spoken or written, of the matters concerning a particular topic. This could refer to anything from a courtroom testimony to a grade schooler’s book report .
Really, when people talk about “reports,” they’re usually referring to official documents outlining the facts of a topic, typically written by an expert on the subject or someone assigned to investigate it. There are different types of reports, explained in the next section, but they mostly fit this description.
What kind of information is shared in reports? Although all facts are welcome, reports, in particular, tend to feature these types of content:
- Details of an event or situation
- The consequences or ongoing effect of an event or situation
- Evaluation of statistical data or analytics
- Interpretations from the information in the report
- Predictions or recommendations based on the information in the report
- How the information relates to other events or reports
Reports are closely related to essay writing , although there are some clear distinctions. While both rely on facts, essays add the personal opinions and arguments of the authors. Reports typically stick only to the facts, although they may include some of the author’s interpretation of these facts, most likely in the conclusion.
Moreover, reports are heavily organized, commonly with tables of contents and copious headings and subheadings. This makes it easier for readers to scan reports for the information they’re looking for. Essays, on the other hand, are meant to be read start to finish, not browsed for specific insights.
There are a few different types of reports, depending on the purpose and to whom you present your report. Here’s a quick list of the common types of reports:
- Academic report: Tests a student’s comprehension of the subject matter, such as book reports, reports on historical events, and biographies
- Business reports: Identifies information useful in business strategy, such as marketing reports, internal memos, SWOT analysis, and feasibility reports
- Scientific reports: Shares research findings, such as research papers and case studies, typically in science journals
Reports can be further divided into categories based on how they are written. For example, a report could be formal or informal, short or long, and internal or external. In business, a vertical report shares information with people on different levels of the hierarchy (i.e., people who work above you and below you), while a lateral report is for people on the author’s same level, but in different departments.
There are as many types of reports as there are writing styles, but in this guide, we focus on academic reports, which tend to be formal and informational.
>>Read More: What Is Academic Writing?
The report format depends on the type of report and the requirements of the assignment. While reports can use their own unique structure, most follow this basic template:
- Executive summary: Just like an abstract in an academic paper, an executive summary is a standalone section that summarizes the findings in your report so readers know what to expect. These are mostly for official reports and less so for school reports.
- Introduction: Setting up the body of the report, your introduction explains the overall topic that you’re about to discuss, with your thesis statement and any need-to-know background information before you get into your own findings.
- Body: The body of the report explains all your major discoveries, broken up into headings and subheadings. The body makes up the majority of the entire report; whereas the introduction and conclusion are just a few paragraphs each, the body can go on for pages.
- Conclusion: The conclusion is where you bring together all the information in your report and come to a definitive interpretation or judgment. This is usually where the author inputs their own personal opinions or inferences.
If you’re familiar with how to write a research paper , you’ll notice that report writing follows the same introduction-body-conclusion structure, sometimes adding an executive summary. Reports usually have their own additional requirements as well, such as title pages and tables of content, which we explain in the next section.
There are no firm requirements for what’s included in a report. Every school, company, laboratory, task manager, and teacher can make their own format, depending on their unique needs. In general, though, be on the lookout for these particular requirements—they tend to crop up a lot:
- Title page: Official reports often use a title page to keep things organized; if a person has to read multiple reports, title pages make them easier to keep track of.
- Table of contents: Just like in books, the table of contents helps readers go directly to the section they’re interested in, allowing for faster browsing.
- Page numbering: A common courtesy if you’re writing a longer report, page numbering makes sure the pages are in order in the case of mix-ups or misprints.
- Headings and subheadings: Reports are typically broken up into sections, divided by headings and subheadings, to facilitate browsing and scanning.
- Citations: If you’re citing information from another source, the citations guidelines tell you the recommended format.
- Works cited page: A bibliography at the end of the report lists credits and the legal information for the other sources you got information from.
As always, refer to the assignment for the specific guidelines on each of these. The people who read the report should tell you which style guides or formatting they require.
Now let’s get into the specifics of how to write a report. Follow the seven steps on report writing below to take you from an idea to a completed paper.
1 Choose a topic based on the assignment
Before you start writing, you need to pick the topic of your report. Often, the topic is assigned for you, as with most business reports, or predetermined by the nature of your work, as with scientific reports. If that’s the case, you can ignore this step and move on.
If you’re in charge of choosing your own topic, as with a lot of academic reports, then this is one of the most important steps in the whole writing process. Try to pick a topic that fits these two criteria:
- There’s adequate information: Choose a topic that’s not too general but not too specific, with enough information to fill your report without padding, but not too much that you can’t cover everything.
- It’s something you’re interested in: Although this isn’t a strict requirement, it does help the quality of a report if you’re engaged by the subject matter.
Of course, don’t forget the instructions of the assignment, including length, so keep those in the back of your head when deciding.
2 Conduct research
With business and scientific reports, the research is usually your own or provided by the company—although there’s still plenty of digging for external sources in both.
For academic papers, you’re largely on your own for research, unless you’re required to use class materials. That’s one of the reasons why choosing the right topic is so crucial; you won’t go far if the topic you picked doesn’t have enough available research.
The key is to search only for reputable sources: official documents, other reports, research papers, case studies, books from respected authors, etc. Feel free to use research cited in other similar reports. You can often find a lot of information online through search engines, but a quick trip to the library can also help in a pinch.
3 Write a thesis statement
Before you go any further, write a thesis statement to help you conceptualize the main theme of your report. Just like the topic sentence of a paragraph, the thesis statement summarizes the main point of your writing, in this case, the report.
Once you’ve collected enough research, you should notice some trends and patterns in the information. If these patterns all infer or lead up to a bigger, overarching point, that’s your thesis statement.
For example, if you were writing a report on the wages of fast-food employees, your thesis might be something like, “Although wages used to be commensurate with living expenses, after years of stagnation they are no longer adequate.” From there, the rest of your report will elaborate on that thesis, with ample evidence and supporting arguments.
It’s good to include your thesis statement in both the executive summary and introduction of your report, but you still want to figure it out early so you know which direction to go when you work on your outline next.
4 Prepare an outline
Writing an outline is recommended for all kinds of writing, but it’s especially useful for reports given their emphasis on organization. Because reports are often separated by headings and subheadings, a solid outline makes sure you stay on track while writing without missing anything.
Really, you should start thinking about your outline during the research phase, when you start to notice patterns and trends. If you’re stuck, try making a list of all the key points, details, and evidence you want to mention. See if you can fit them into general and specific categories, which you can turn into headings and subheadings respectively.
5 Write a rough draft
Actually writing the rough draft , or first draft, is usually the most time-consuming step. Here’s where you take all the information from your research and put it into words. To avoid getting overwhelmed, simply follow your outline step by step to make sure you don’t accidentally leave out anything.
Don’t be afraid to make mistakes; that’s the number one rule for writing a rough draft. Expecting your first draft to be perfect adds a lot of pressure. Instead, write in a natural and relaxed way, and worry about the specific details like word choice and correcting mistakes later. That’s what the last two steps are for, anyway.
6 Revise and edit your report
Once your rough draft is finished, it’s time to go back and start fixing the mistakes you ignored the first time around. (Before you dive right back in, though, it helps to sleep on it to start editing fresh, or at least take a small break to unwind from writing the rough draft.)
We recommend first rereading your report for any major issues, such as cutting or moving around entire sentences and paragraphs. Sometimes you’ll find your data doesn’t line up, or that you misinterpreted a key piece of evidence. This is the right time to fix the “big picture” mistakes and rewrite any longer sections as needed.
If you’re unfamiliar with what to look for when editing, you can read our previous guide with some more advanced self-editing tips .
7 Proofread and check for mistakes
Last, it pays to go over your report one final time, just to optimize your wording and check for grammatical or spelling mistakes. In the previous step you checked for “big picture” mistakes, but here you’re looking for specific, even nitpicky problems.
A writing assistant like Grammarly flags those issues for you. Grammarly’s free version points out any spelling and grammatical mistakes while you write, with suggestions to improve your writing that you can apply with just one click. The Premium version offers even more advanced features, such as tone adjustments and word choice recommendations for taking your writing to the next level.

- Search Search Please fill out this field.
What Is a Research Report?
Understanding research reports, financial analyst research reports, research report impact, conflicts of interest.
- Fundamental Analysis
What Is a Research Report? How They're Produced and Impact
James Chen, CMT is an expert trader, investment adviser, and global market strategist.
:max_bytes(150000):strip_icc():format(webp)/photo__james_chen-5bfc26144cedfd0026c00af8.jpeg)
A research report is a document prepared by an analyst or strategist who is a part of the investment research team in a stock brokerage or investment bank . A research report may focus on a specific stock or industry sector, a currency, commodity or fixed-income instrument, or on a geographic region or country. Research reports generally, but not always, have actionable recommendations such as investment ideas that investors can act upon.
Research reports are produced by a variety of sources, ranging from market research firms to in-house departments at large organizations. When applied to the investment industry, the term usually refers to sell-side research, or investment research produced by brokerage houses.
Such research is disseminated to the institutional and retail clients of the brokerage that produces it. Research produced by the buy-side, which includes pension funds, mutual funds, and portfolio managers , is usually for internal use only and is not distributed to external parties.
Financial analysts may produce research reports for the purpose of supporting a particular recommendation, such as whether to buy or sell a particular security or whether a client should consider a particular financial product. For example, an analyst may create a report in regards to a new offering being proposed by a company. The report could include relevant metrics regarding the company itself, such as the number of years they have been in operation as well as the names of key stakeholders , along with statistics regarding the current state of the market in which the company participates. Information regarding overall profitability and the intended use of the funds can also be included.
Enthusiasts of the Efficient Market Hypothesis (EMH) might insist that the value of professional analysts' research reports is suspect and that investors likely place too much confidence in the conclusions such analysts make. While a definitive conclusion about this topic is difficult to make because comparisons are not exact, some research papers do exist which claim empirical evidence supporting the value of such reports.
One such paper studied the market for India-based investments and analysts who cover them. The paper was published in the March 2014 edition of the International Research Journal of Business and Management. Its authors concluded that analyst recommendations do have an impact and are beneficial to investors at least in short-term decisions.
While some analysts are functionally unaffiliated, others may be directly or indirectly affiliated with the companies for which they produce reports. Unaffiliated analysts traditionally perform independent research to determine an appropriate recommendation and may have a limited concern regarding the outcome.
Affiliated analysts may feel best served by ensuring any research reports portray clients in a favorable light. Additionally, if an analyst is also an investor in the company on which the report is based, he may have a personal incentive to avoid topics that may result in a lowered valuation of the securities in which he has invested.
:max_bytes(150000):strip_icc():format(webp)/GettyImages-1206249788-47ee99bf82264d57a538463db4e1951e.jpg)
- Terms of Service
- Editorial Policy
- Privacy Policy
- Your Privacy Choices
Organizing Your Social Sciences Research Assignments
- Annotated Bibliography
- Analyzing a Scholarly Journal Article
- Group Presentations
- Dealing with Nervousness
- Using Visual Aids
- Grading Someone Else's Paper
- Types of Structured Group Activities
- Group Project Survival Skills
- Leading a Class Discussion
- Multiple Book Review Essay
- Reviewing Collected Works
- Writing a Case Analysis Paper
- Writing a Case Study
- About Informed Consent
- Writing Field Notes
- Writing a Policy Memo
- Writing a Reflective Paper
- Writing a Research Proposal
- Generative AI and Writing
- Acknowledgments
The purpose of a field report in the social sciences is to describe the deliberate observation of people, places, and/or events and to analyze what has been observed in order to identify and categorize common themes in relation to the research problem underpinning the study. The content represents the researcher's interpretation of meaning found in data that has been gathered during one or more observational events.
Flick, Uwe. The SAGE Handbook of Qualitative Data Collection . London: SAGE Publications, 2018; Lofland, John, David Snow, Leon Anderson, and Lyn H. Lofland. Analyzing Social Settings: A Guide to Qualitative Observation and Analysis. Long Grove, IL: Waveland Press, 2022; Baker, Lynda. "Observation: A Complex Research Method." Library Trends 55 (Summer 2006): 171-189.; Kellehear, Allan. The Unobtrusive Researcher: A Guide to Methods . New York: Routledge, 2020.
How to Approach Writing a Field Report
How to Begin
Field reports are most often assigned in disciplines of the applied social sciences [e.g., social work, anthropology, gerontology, criminal justice, education, law, the health care services] where it is important to build a bridge of relevancy between the theoretical concepts learned in the classroom and the practice of actually doing the work you are being taught to do. Field reports are also common in certain science disciplines [e.g., geology] but these reports are organized differently and serve a different purpose than what is described below.
Professors will assign a field report with the intention of improving your understanding of key theoretical concepts by applying methods of careful and structured observation of, and reflection about, people, places, or phenomena existing in their natural settings. Field reports facilitate the development of data collection techniques and observation skills and they help you to understand how theory applies to real world situations. Field reports are also an opportunity to obtain evidence through methods of observing professional practice that contribute to or challenge existing theories.
We are all observers of people, their interactions, places, and events; however, your responsibility when writing a field report is to conduct research based on data generated by the act of designing a specific study, deliberate observation, synthesis of key findings, and interpretation of their meaning.
When writing a field report you need to:
- Systematically observe and accurately record the varying aspects of a situation . Always approach your field study with a detailed protocol about what you will observe, where you should conduct your observations, and the method by which you will collect and record your data.
- Continuously analyze your observations . Always look for the meaning underlying the actions you observe. Ask yourself: What's going on here? What does this observed activity mean? What else does this relate to? Note that this is an on-going process of reflection and analysis taking place for the duration of your field research.
- Keep the report’s aims in mind while you are observing . Recording what you observe should not be done randomly or haphazardly; you must be focused and pay attention to details. Enter the observation site [i.e., "field"] with a clear plan about what you are intending to observe and record in relation to the research problem while, at the same time, being prepared to adapt to changing circumstances as they may arise.
- Consciously observe, record, and analyze what you hear and see in the context of a theoretical framework . This is what separates data gatherings from reporting. The theoretical framework guiding your field research should determine what, when, and how you observe and act as the foundation from which you interpret your findings in relation to the underlying assumptions embedded in the theoretical framework .
Techniques to Record Your Observations Although there is no limit to the type of data gathering techniques you can use, these are the most frequently used methods:
Note Taking This is the most common and easiest method of recording your observations. Tips for taking notes include: organizing some shorthand symbols beforehand so that recording basic or repeated actions does not impede your ability to observe, using many small paragraphs, which reflect changes in activities, who is talking, etc., and, leaving space on the page so you can write down additional thoughts and ideas about what’s being observed, any theoretical insights, and notes to yourself that are set aside for further investigation. See drop-down tab for additional information about note-taking.
Photography With the advent of smart phones, an almost unlimited number of high quality photographs can be taken of the objects, events, and people observed during a field study. Photographs can help capture an important moment in time as well as document details about the space where your observation takes place. Taking a photograph can save you time in documenting the details of a space that would otherwise require extensive note taking. However, be aware that flash photography could undermine your ability to observe unobtrusively so assess the lighting in your observation space; if it's too dark, you may need to rely on taking notes. Also, you should reject the idea that photographs represent some sort of "window into the world" because this assumption creates the risk of over-interpreting what they show. As with any product of data gathering, you are the sole instrument of interpretation and meaning-making, not the object itself. Video and Audio Recordings Video or audio recording your observations has the positive effect of giving you an unfiltered record of the observation event. It also facilitates repeated analysis of your observations. This can be particularly helpful as you gather additional information or insights during your research. However, these techniques have the negative effect of increasing how intrusive you are as an observer and will often not be practical or even allowed under certain circumstances [e.g., interaction between a doctor and a patient] and in certain organizational settings [e.g., a courtroom]. Illustrations/Drawings This does not refer to an artistic endeavor but, rather, refers to the possible need, for example, to draw a map of the observation setting or illustrating objects in relation to people's behavior. This can also take the form of rough tables, charts, or graphs documenting the frequency and type of activities observed. These can be subsequently placed in a more readable format when you write your field report. To save time, draft a table [i.e., columns and rows] on a separate piece of paper before an observation if you know you will be entering data in that way.
NOTE: You may consider using a laptop or other electronic device to record your notes as you observe, but keep in mind the possibility that the clicking of keys while you type or noises from your device can be obtrusive, whereas writing your notes on paper is relatively quiet and unobtrusive. Always assess your presence in the setting where you're gathering the data so as to minimize your impact on the subject or phenomenon being studied.
ANOTHER NOTE: Techniques of deliberate observation and data gathering are not innate skills; they are skills that must be learned and practiced in order to achieve proficiency. Before your first observation, practice the technique you plan to use in a setting similar to your study site [e.g., take notes about how people choose to enter checkout lines at a grocery store if your research involves examining the choice patterns of unrelated people forced to queue in busy social settings]. When the act of data gathering counts, you'll be glad you practiced beforehand.
YET ANOTHER NOTE: An issue rarely discussed in the literature about conducting field research is whether you should move around the study site while observing or remaining situated in one place. Moving around can be intrusive, but it facilitates observing people's behavior from multiple vectors. However, if you remain in one place throughout the observation [or during each observation], you will eventually blend into the background and diminish the chance of unintentionally influencing people's behavior. If the site has a complex set of interactions or interdependent activities [e.g., a play ground], consider moving around; if the study site is relatively fixed [e.g., a classroom], then consider staying in one place while observing.
Examples of Things to Document While Observing
- Physical setting . The characteristics of an occupied space and the human use of the place where the observation(s) are being conducted.
- Objects and material culture . This refers to the presence, placement, and arrangement of objects that impact the behavior or actions of those being observed. If applicable, describe the cultural artifacts representing the beliefs [i.e., the values, ideas, attitudes, and assumptions] of the individuals you are observing [e.g., the choice of particular types of clothing in the observation of family gatherings during culturally specific holidays].
- Use of language . Don't just observe but listen to what is being said, how is it being said, and the tone of conversations among participants.
- Behavior cycles . This refers to documenting when and who performs what behavior or task and how often they occur. Record at which stage this behavior is occurring within the setting.
- The order in which events unfold . Note sequential patterns of behavior or the moment when actions or events take place and their significance. Also, be prepared to note moments that diverge from these sequential patterns of behavior or actions.
- Physical characteristics of subjects. If relevant, document personal characteristics of individuals being observed. Note that, unless this data can be verified in interviews or from documentary evidence, you should only focus on characteristics that can be clearly observed [e.g., clothing, physical appearance, body language].
- Expressive body movements . This would include things like body posture or facial expressions. Note that it may be relevant to also assess whether expressive body movements support or contradict the language used in conversation [e.g., detecting sarcasm].
Brief notes about all of these examples contextualize your observations; however, your observation notes will be guided primarily by your theoretical framework, keeping in mind that your observations will feed into and potentially modify or alter these frameworks.
Sampling Techniques
Sampling refers to the process used to select a portion of the population for study . Qualitative research, of which observation is one method of data gathering, is generally based on non-probability and purposive sampling rather than probability or random approaches characteristic of quantitatively-driven studies. Sampling in observational research is flexible and often continues until no new themes emerge from the data, a point referred to as data saturation.
All sampling decisions are made for the explicit purpose of obtaining the richest possible source of information to answer the research questions. Decisions about sampling assumes you know what you want to observe, what behaviors are important to record, and what research problem you are addressing before you begin the study. These questions determine what sampling technique you should use, so be sure you have adequately answered them before selecting a sampling method.
Ways to sample when conducting an observation include:
- Ad Libitum Sampling -- this approach is not that different from what people do at the zoo; they observe whatever seems interesting at the moment. There is no organized system of recording the observations; you just note whatever seems relevant at the time. The advantage of this method is that you are often able to observe relatively rare or unusual behaviors that might be missed by more deliberately designed sampling methods. This method is also useful for obtaining preliminary observations that can be used to develop your final field study. Problems using this method include the possibility of inherent bias toward conspicuous behaviors or individuals, thereby missing mundane or repeated patterns of behavior, and that you may miss brief interactions in social settings.
- Behavior Sampling -- this involves watching the entire group of subjects and recording each occurrence of a specific behavior of interest and with reference to which individuals were involved. The method is useful in recording rare behaviors missed by other sampling methods and is often used in conjunction with focal or scan methods [see below]. However, sampling can be biased towards particular conspicuous behaviors.
- Continuous Recording -- provides a faithful record of behavior including frequencies, durations, and latencies [the time that elapses between a stimulus and the response to it]. This is a very demanding method because you are trying to record everything within the setting and, thus, measuring reliability may be sacrificed. In addition, durations and latencies are only reliable if subjects remain present throughout the collection of data. However, this method facilitates analyzing sequences of behaviors and ensures obtaining a wealth of data about the observation site and the people within it. The use of audio or video recording is most useful with this type of sampling.
- Focal Sampling -- this involves observing one individual for a specified amount of time and recording all instances of that individual's behavior. Usually you have a set of predetermined categories or types of behaviors that you are interested in observing [e.g., when a teacher walks around the classroom] and you keep track of the duration of those behaviors. This approach doesn't tend to bias one behavior over another and provides significant detail about a individual's behavior. However, with this method, you likely have to conduct a lot of focal samples before you have a good idea about how group members interact. It can also be difficult within certain settings to keep one individual in sight for the entire period of the observation without being intrusive.
- Instantaneous Sampling -- this is where observation sessions are divided into short intervals divided by sample points. At each sample point the observer records if predetermined behaviors of interest are taking place. This method is not effective for recording discrete events of short duration and, frequently, observers will want to record novel behaviors that occur slightly before or after the point of sampling, creating a sampling error. Though not exact, this method does give you an idea of durations and is relatively easy to do. It is also good for recording behavior patterns occurring at a specific instant, such as, movement or body positions.
- One-Zero Sampling -- this is very similar to instantaneous sampling, only the observer records if the behaviors of interest have occurred at any time during an interval instead of at the instant of the sampling point. The method is useful for capturing data on behavior patterns that start and stop repeatedly and rapidly, but that last only for a brief period of time. The disadvantage of this approach is that you get a dimensionless score for an entire recording session, so you only get one one data point for each recording session.
- Scan Sampling -- this method involves taking a census of the entire observed group at predetermined time periods and recording what each individual is doing at that moment. This is useful for obtaining group behavioral data and allows for data that are evenly representative across individuals and periods of time. On the other hand, this method may be biased towards more conspicuous behaviors and you may miss a lot of what is going on between observations, especially rare or unusual behaviors. It is also difficult to record more than a few individuals in a group setting without missing what each individual is doing at each predetermined moment in time [e.g., children sitting at a table during lunch at school]. The use of audio or video recording is useful with this type of sampling.
Alderks, Peter. Data Collection. Psychology 330 Course Documents. Animal Behavior Lab. University of Washington; Emerson, Robert M. Contemporary Field Research: Perspectives and Formulations . 2nd ed. Prospect Heights, IL: Waveland Press, 2001; Emerson, Robert M. et al. “Participant Observation and Fieldnotes.” In Handbook of Ethnography . Paul Atkinson et al., eds. (Thousand Oaks, CA: Sage, 2001), 352-368; Emerson, Robert M. et al. Writing Ethnographic Fieldnotes . 2nd ed. Chicago, IL: University of Chicago Press, 2011; Ethnography, Observational Research, and Narrative Inquiry. Writing@CSU. Colorado State University; Hazel, Spencer. "The Paradox from Within: Research Participants Doing-Being-Observed." Qualitative Research 16 (August 2016): 446-457; Pace, Tonio. Writing Field Reports. Scribd Online Library; Presser, Jon and Dona Schwartz. “Photographs within the Sociological Research Process.” In Image-based Research: A Sourcebook for Qualitative Researchers . Jon Prosser, editor (London: Falmer Press, 1998), pp. 115-130; Pyrczak, Fred and Randall R. Bruce. Writing Empirical Research Reports: A Basic Guide for Students of the Social and Behavioral Sciences . 5th ed. Glendale, CA: Pyrczak Publishing, 2005; Report Writing. UniLearning. University of Wollongong, Australia; Wolfinger, Nicholas H. "On Writing Fieldnotes: Collection Strategies and Background Expectancies.” Qualitative Research 2 (April 2002): 85-95; Writing Reports. Anonymous. The Higher Education Academy.
Structure and Writing Style
How you choose to format your field report is determined by the research problem, the theoretical framework that is driving your analysis, the observations that you make, and/or specific guidelines established by your professor. Since field reports do not have a standard format, it is worthwhile to determine from your professor what the preferred structure and organization should be before you begin to write. Note that field reports should be written in the past tense. With this in mind, most field reports in the social sciences include the following elements:
I. Introduction The introduction should describe the research problem, the specific objectives of your research, and the important theories or concepts underpinning your field study. The introduction should describe the nature of the organization or setting where you are conducting the observation, what type of observations you have conducted, what your focus was, when you observed, and the methods you used for collecting the data. Collectively, this descriptive information should support reasons why you chose the observation site and the people or events within it. You should also include a review of pertinent literature related to the research problem, particularly if similar methods were used in prior studies. Conclude your introduction with a statement about how the rest of the paper is organized.
II. Description of Activities
Your readers only knowledge and understanding of what happened will come from the description section of your report because they were not witnesses to the situation, people, or events that you are writing about. Given this, it is crucial that you provide sufficient details to place the analysis that will follow into proper context; don't make the mistake of providing a description without context. The description section of a field report is similar to a well written piece of journalism. Therefore, a useful approach to systematically describing the varying aspects of an observed situation is to answer the "Five W’s of Investigative Reporting." As Dubbels notes [p. 19], these are:
- What -- describe what you observed. Note the temporal, physical, and social boundaries you imposed to limit the observations you made. What were your general impressions of the situation you were observing. For example, as a student teacher, what is your impression of the application of iPads as a learning device in a history class; as a cultural anthropologist, what is your impression of women's participation in a Native American religious ritual?
- Where -- provide background information about the setting of your observation and, if necessary, note important material objects that are present that help contextualize the observation [e.g., arrangement of computers in relation to student engagement with the teacher].
- When -- record factual data about the day and the beginning and ending time of each observation. Note that it may also be necessary to include background information or key events which impact upon the situation you were observing [e.g., observing the ability of teachers to re-engage students after coming back from an unannounced fire drill].
- Who -- note background and demographic information about the individuals being observed e.g., age, gender, ethnicity, and/or any other variables relevant to your study]. Record who is doing what and saying what, as well as, who is not doing or saying what. If relevant, be sure to record who was missing from the observation.
- Why -- why were you doing this? Describe the reasons for selecting particular situations to observe. Note why something happened. Also note why you may have included or excluded certain information.
III. Interpretation and Analysis
Always place the analysis and interpretations of your field observations within the larger context of the theoretical assumptions and issues you described in the introduction. Part of your responsibility in analyzing the data is to determine which observations are worthy of comment and interpretation, and which observations are more general in nature. It is your theoretical framework that allows you to make these decisions. You need to demonstrate to the reader that you are conducting the field work through the eyes of an informed viewer and from the perspective of a casual observer.
Here are some questions to ask yourself when analyzing your observations:
- What is the meaning of what you have observed?
- Why do you think what you observed happened? What evidence do you have for your reasoning?
- What events or behaviors were typical or widespread? If appropriate, what was unusual or out of the ordinary? How were they distributed among categories of people?
- Do you see any connections or patterns in what you observed?
- Why did the people you observed proceed with an action in the way that they did? What are the implications of this?
- Did the stated or implicit objectives of what you were observing match what was achieved?
- What were the relative merits of the behaviors you observed?
- What were the strengths and weaknesses of the observations you recorded?
- Do you see connections between what you observed and the findings of similar studies identified from your review of the literature?
- How do your observations fit into the larger context of professional practice? In what ways have your observations possibly changed or affirmed your perceptions of professional practice?
- Have you learned anything from what you observed?
NOTE: Only base your interpretations on what you have actually observed. Do not speculate or manipulate your observational data to fit into your study's theoretical framework.
IV. Conclusion and Recommendations
The conclusion should briefly recap of the entire study, reiterating the importance or significance of your observations. Avoid including any new information. You should also state any recommendations you may have based on the results of your study. Be sure to describe any unanticipated problems you encountered and note the limitations of your study. The conclusion should not be more than two or three paragraphs.
V. Appendix
This is where you would place information that is not essential to explaining your findings, but that supports your analysis [especially repetitive or lengthy information], that validates your conclusions, or that contextualizes a related point that helps the reader understand the overall report. Examples of information that could be included in an appendix are figures/tables/charts/graphs of results, statistics, pictures, maps, drawings, or, if applicable, transcripts of interviews. There is no limit to what can be included in the appendix or its format [e.g., a DVD recording of the observation site], provided that it is relevant to the study's purpose and reference is made to it in the report. If information is placed in more than one appendix ["appendices"], the order in which they are organized is dictated by the order they were first mentioned in the text of the report.
VI. References
List all sources that you consulted and obtained information from while writing your field report. Note that field reports generally do not include further readings or an extended bibliography. However, consult with your professor concerning what your list of sources should be included and be sure to write them in the preferred citation style of your discipline or is preferred by your professor [i.e., APA, Chicago, MLA, etc.].
Alderks, Peter. Data Collection. Psychology 330 Course Documents. Animal Behavior Lab. University of Washington; Dubbels, Brock R. Exploring the Cognitive, Social, Cultural, and Psychological Aspects of Gaming and Simulations . Hershey, PA: IGI Global, 2018; Emerson, Robert M. Contemporary Field Research: Perspectives and Formulations . 2nd ed. Prospect Heights, IL: Waveland Press, 2001; Emerson, Robert M. et al. “Participant Observation and Fieldnotes.” In Handbook of Ethnography . Paul Atkinson et al., eds. (Thousand Oaks, CA: Sage, 2001), 352-368; Emerson, Robert M. et al. Writing Ethnographic Fieldnotes . 2nd ed. Chicago, IL: University of Chicago Press, 2011; Ethnography, Observational Research, and Narrative Inquiry. Writing@CSU. Colorado State University; Pace, Tonio. Writing Field Reports. Scribd Online Library; Pyrczak, Fred and Randall R. Bruce. Writing Empirical Research Reports: A Basic Guide for Students of the Social and Behavioral Sciences . 5th ed. Glendale, CA: Pyrczak Publishing, 2005; Report Writing. UniLearning. University of Wollongong, Australia; Wolfinger, Nicholas H. "On Writing Fieldnotes: Collection Strategies and Background Expectancies.” Qualitative Research 2 (April 2002): 85-95; Writing Reports. Anonymous. The Higher Education Academy.
- << Previous: Writing a Case Study
- Next: About Informed Consent >>
- Last Updated: Jun 3, 2024 9:44 AM
- URL: https://libguides.usc.edu/writingguide/assignments
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, generate accurate citations for free.
- Knowledge Base
- Research paper
Writing a Research Paper Conclusion | Step-by-Step Guide
Published on October 30, 2022 by Jack Caulfield . Revised on April 13, 2023.
- Restate the problem statement addressed in the paper
- Summarize your overall arguments or findings
- Suggest the key takeaways from your paper
The content of the conclusion varies depending on whether your paper presents the results of original empirical research or constructs an argument through engagement with sources .
Instantly correct all language mistakes in your text
Upload your document to correct all your mistakes in minutes
Table of contents
Step 1: restate the problem, step 2: sum up the paper, step 3: discuss the implications, research paper conclusion examples, frequently asked questions about research paper conclusions.
The first task of your conclusion is to remind the reader of your research problem . You will have discussed this problem in depth throughout the body, but now the point is to zoom back out from the details to the bigger picture.
While you are restating a problem you’ve already introduced, you should avoid phrasing it identically to how it appeared in the introduction . Ideally, you’ll find a novel way to circle back to the problem from the more detailed ideas discussed in the body.
For example, an argumentative paper advocating new measures to reduce the environmental impact of agriculture might restate its problem as follows:
Meanwhile, an empirical paper studying the relationship of Instagram use with body image issues might present its problem like this:
“In conclusion …”
Avoid starting your conclusion with phrases like “In conclusion” or “To conclude,” as this can come across as too obvious and make your writing seem unsophisticated. The content and placement of your conclusion should make its function clear without the need for additional signposting.
Don't submit your assignments before you do this
The academic proofreading tool has been trained on 1000s of academic texts. Making it the most accurate and reliable proofreading tool for students. Free citation check included.
Try for free
Having zoomed back in on the problem, it’s time to summarize how the body of the paper went about addressing it, and what conclusions this approach led to.
Depending on the nature of your research paper, this might mean restating your thesis and arguments, or summarizing your overall findings.
Argumentative paper: Restate your thesis and arguments
In an argumentative paper, you will have presented a thesis statement in your introduction, expressing the overall claim your paper argues for. In the conclusion, you should restate the thesis and show how it has been developed through the body of the paper.
Briefly summarize the key arguments made in the body, showing how each of them contributes to proving your thesis. You may also mention any counterarguments you addressed, emphasizing why your thesis holds up against them, particularly if your argument is a controversial one.
Don’t go into the details of your evidence or present new ideas; focus on outlining in broad strokes the argument you have made.
Empirical paper: Summarize your findings
In an empirical paper, this is the time to summarize your key findings. Don’t go into great detail here (you will have presented your in-depth results and discussion already), but do clearly express the answers to the research questions you investigated.
Describe your main findings, even if they weren’t necessarily the ones you expected or hoped for, and explain the overall conclusion they led you to.
Having summed up your key arguments or findings, the conclusion ends by considering the broader implications of your research. This means expressing the key takeaways, practical or theoretical, from your paper—often in the form of a call for action or suggestions for future research.
Argumentative paper: Strong closing statement
An argumentative paper generally ends with a strong closing statement. In the case of a practical argument, make a call for action: What actions do you think should be taken by the people or organizations concerned in response to your argument?
If your topic is more theoretical and unsuitable for a call for action, your closing statement should express the significance of your argument—for example, in proposing a new understanding of a topic or laying the groundwork for future research.
Empirical paper: Future research directions
In a more empirical paper, you can close by either making recommendations for practice (for example, in clinical or policy papers), or suggesting directions for future research.
Whatever the scope of your own research, there will always be room for further investigation of related topics, and you’ll often discover new questions and problems during the research process .
Finish your paper on a forward-looking note by suggesting how you or other researchers might build on this topic in the future and address any limitations of the current paper.
Full examples of research paper conclusions are shown in the tabs below: one for an argumentative paper, the other for an empirical paper.
- Argumentative paper
- Empirical paper
While the role of cattle in climate change is by now common knowledge, countries like the Netherlands continually fail to confront this issue with the urgency it deserves. The evidence is clear: To create a truly futureproof agricultural sector, Dutch farmers must be incentivized to transition from livestock farming to sustainable vegetable farming. As well as dramatically lowering emissions, plant-based agriculture, if approached in the right way, can produce more food with less land, providing opportunities for nature regeneration areas that will themselves contribute to climate targets. Although this approach would have economic ramifications, from a long-term perspective, it would represent a significant step towards a more sustainable and resilient national economy. Transitioning to sustainable vegetable farming will make the Netherlands greener and healthier, setting an example for other European governments. Farmers, policymakers, and consumers must focus on the future, not just on their own short-term interests, and work to implement this transition now.
As social media becomes increasingly central to young people’s everyday lives, it is important to understand how different platforms affect their developing self-conception. By testing the effect of daily Instagram use among teenage girls, this study established that highly visual social media does indeed have a significant effect on body image concerns, with a strong correlation between the amount of time spent on the platform and participants’ self-reported dissatisfaction with their appearance. However, the strength of this effect was moderated by pre-test self-esteem ratings: Participants with higher self-esteem were less likely to experience an increase in body image concerns after using Instagram. This suggests that, while Instagram does impact body image, it is also important to consider the wider social and psychological context in which this usage occurs: Teenagers who are already predisposed to self-esteem issues may be at greater risk of experiencing negative effects. Future research into Instagram and other highly visual social media should focus on establishing a clearer picture of how self-esteem and related constructs influence young people’s experiences of these platforms. Furthermore, while this experiment measured Instagram usage in terms of time spent on the platform, observational studies are required to gain more insight into different patterns of usage—to investigate, for instance, whether active posting is associated with different effects than passive consumption of social media content.
If you’re unsure about the conclusion, it can be helpful to ask a friend or fellow student to read your conclusion and summarize the main takeaways.
- Do they understand from your conclusion what your research was about?
- Are they able to summarize the implications of your findings?
- Can they answer your research question based on your conclusion?
You can also get an expert to proofread and feedback your paper with a paper editing service .
The conclusion of a research paper has several key elements you should make sure to include:
- A restatement of the research problem
- A summary of your key arguments and/or findings
- A short discussion of the implications of your research
No, it’s not appropriate to present new arguments or evidence in the conclusion . While you might be tempted to save a striking argument for last, research papers follow a more formal structure than this.
All your findings and arguments should be presented in the body of the text (more specifically in the results and discussion sections if you are following a scientific structure). The conclusion is meant to summarize and reflect on the evidence and arguments you have already presented, not introduce new ones.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the “Cite this Scribbr article” button to automatically add the citation to our free Citation Generator.
Caulfield, J. (2023, April 13). Writing a Research Paper Conclusion | Step-by-Step Guide. Scribbr. Retrieved August 12, 2024, from https://www.scribbr.com/research-paper/research-paper-conclusion/
Is this article helpful?

Jack Caulfield
Other students also liked, writing a research paper introduction | step-by-step guide, how to create a structured research paper outline | example, checklist: writing a great research paper, what is your plagiarism score.
Hindenburg alleges India market regulator chief held stake in offshore funds used by Adani Group
- Medium Text

- Adani Enterprises Ltd Follow
- Adani Group Follow
- Hindenburg Research LLC Follow
Sign up here.
Reporting by Jayshree.P. Upadhyay; Additional reporting by Ira Dugal, Neha Arora and Jahnavi Nidumolu; Editing by Jacqueline Wong, William Mallard, David Goodman and Toby Chopra
Our Standards: The Thomson Reuters Trust Principles. , opens new tab

US Senator Graham calls Ukraine's incursion into Russia 'bold', 'beautiful'
U.S. Republican Senator Lindsey Graham called Ukraine's shock cross-border operation into Russia's Kursk region "brilliant" and "bold" in a visit to Kyiv on Monday, and he urged the Biden administration to provide Ukraine with the weapons it needs.

Writing up a Research Report
- First Online: 10 November 2021
Cite this chapter

- Stefan Hunziker 3 &
- Michael Blankenagel 3
3399 Accesses
A research report is one big argument how and why you came up with your conclusions. To make it a convincing argument, a typical guiding structure has developed. In the different chapters, distinct issues need to be addressed to explain to the reader why your conclusions are valid. The governing principle for writing the report is full disclosure: to explain everything and ensure replicability by another researcher.
This is a preview of subscription content, log in via an institution to check access.
Access this chapter
Subscribe and save.
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
- Available as PDF
- Read on any device
- Instant download
- Own it forever
- Available as EPUB and PDF
Tax calculation will be finalised at checkout
Purchases are for personal use only
Institutional subscriptions
Barros, L. O. (2016). The only academic phrasebook you’ll ever need. Createspace Independent Publishing Platform.
Google Scholar
Field, A. (2016). An adventure in statistics. The reality enigma . SAGE.
Field, A. (2020). Discovering statistics using IBM SPSS statistics (5th ed.). SAGE.
Früh, M., Keimer, I., & Blankenagel, M. (2019). The impact of Balanced Scorecard excellence on shareholder returns. IFZ Working Paper No. 0003/2019. Retrieved June 09, 2021, from https://zenodo.org/record/2571603#.YMDUafkzZaQ .
Yin, R. K. (2013). Case study research: Design and methods (5th ed.). SAGE.
Download references
Author information
Authors and affiliations.
Wirtschaft/IFZ – Campus Zug-Rotkreuz, Hochschule Luzern, Zug-Rotkreuz, Zug , Switzerland
Stefan Hunziker & Michael Blankenagel
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Stefan Hunziker .
Rights and permissions
Reprints and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Fachmedien Wiesbaden GmbH, part of Springer Nature
About this chapter
Hunziker, S., Blankenagel, M. (2021). Writing up a Research Report. In: Research Design in Business and Management. Springer Gabler, Wiesbaden. https://doi.org/10.1007/978-3-658-34357-6_4
Download citation
DOI : https://doi.org/10.1007/978-3-658-34357-6_4
Published : 10 November 2021
Publisher Name : Springer Gabler, Wiesbaden
Print ISBN : 978-3-658-34356-9
Online ISBN : 978-3-658-34357-6
eBook Packages : Business and Economics (German Language)
Share this chapter
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Publish with us
Policies and ethics
- Find a journal
- Track your research
Along with Stanford news and stories, show me:
- Student information
- Faculty/Staff information
We want to provide announcements, events, leadership messages and resources that are relevant to you. Your selection is stored in a browser cookie which you can remove at any time using “Clear all personalization” below.
The Stanford Doerr School of Sustainability has selected eight interconnected Solution Areas to focus its research efforts over the next decade. This new research plan amplifies the school’s ability to translate Stanford research into large-scale solutions and inform key decision makers in policy and business.
Selected based on extensive faculty input and assessment of where Stanford can make the most meaningful impact, the eight areas are: climate; water; energy; food; risk, resilience, and adaptation; nature; cities; and platforms and tools for monitoring and decision making.
“Solution Areas identify and leverage the critical junctions between the most pressing global sustainability challenges and the areas where Stanford has the talent and expertise to find solutions,” said Dean Arun Majumdar. “This collaborative all-campus approach expands and strengthens our commitment to using all the power we have – the knowledge, the education, the talent, the innovation, the resources, the influence – to build a thriving planet for future generations.”
‘Integrative Projects’ and ‘Flagship Destinations’
In each Solution Area, the school plans to build two types of research initiatives. One type, called Integrative Projects, will be managed by the school’s institutes, including the Stanford Woods Institute for the Environment , the Precourt Institute for Energy , and a planned Sustainable Societies Institute.
Integrative Projects will be organized around decade-long research themes and dedicated to creating solutions through interdisciplinary collaboration, engagement with partners beyond Stanford, identifying significant knowledge gaps, and understanding systems.
According to Chris Field , the Perry L. McCarty Director of the Stanford Woods Institute for the Environment and a professor in the Stanford Doerr School of Sustainability and the School of Humanities and Sciences , the new commitment to these areas “will provide both resources and coordination that expand Stanford faculty’s capacity to deliver sustainability solutions at scale.”
A second type of research initiative, called Flagship Destinations, is managed by Stanford’s Sustainability Accelerator . Flagship Destinations are targets for the pace and scale of work to address challenges facing Earth, climate, and society. For example, the school’s first Flagship Destination, announced in 2023 , calls for enabling the removal of billions of tons of planet-warming gases annually from Earth’s atmosphere by the middle of this century. By working backward from sustainability targets in consultation with faculty and external experts, this initiative seeks to rapidly translate Stanford research into policy and technology solutions. Additional Flagship Destinations will be announced later this week.
Whereas Integrative Projects are designed to produce knowledge and evidence that can eventually lead to solutions, Flagship Destination projects are intended to help verify and demonstrate that well-studied solutions can succeed at large scale so they can be launched out of Stanford and implemented for the benefit of humanity and our planet. Scalable solutions nurtured and launched through these projects could take the form of policy frameworks, open-source platforms, nonprofit organizations, new for-profit companies, and ongoing collaborations all committed to addressing pressing sustainability challenges.
“By working together in these Solution Areas across disciplines and with collaborators beyond the university, we maximize our ability to have positive impacts on the timeframe and scale needed for the planet and humanity,” said Scott Fendorf , senior associate dean for integrative initiatives and the Terry Huffington Professor in the Stanford Doerr School of Sustainability.
Workshops will be held with faculty and external experts to develop research strategies for each Solution Area on a rolling basis. Strategy workshops, opportunities to provide input on future Integrative Projects, and requests for proposals (open to all Stanford faculty) will be announced in the coming months.
Related message from leadership: Read a letter to faculty about the new Solution Areas from Dean Majumdar with Precourt Institute for Energy director William Chueh; Stanford Woods Institute for the Environment director Chris Field; Accelerator faculty director Yi Cui and executive director Charlotte Pera; and Integrative Initiatives associate dean Jenna Davis and senior associate dean Scott Fendorf.
Billionaire Gautam Adani’s Fortune Slips Again After Hindenburg Targets Indian Regulator
- Share to Facebook
- Share to Twitter
- Share to Linkedin
Shares of Indian conglomerate Adani Group’s listed firms slumped Monday, after a report by U.S. short seller Hindenburg Group accused the head of India’s market regulator of a conflict of interest that allegedly prevented a proper investigation into Hindenburg’s earlier claims of fraud and stock manipulation by the billionaire-owned conglomerate.
Adani Group chairperson and founder Gautam Adani's fortune slipped again on Monday after ... [+] Hindenburg's latest report.
Shares of the conglomerate’s flagship firm Adani Enterprises dropped more than 5% to $35.9 (Rs 3,013) after Indian stock markets opened on Monday, before recovering to $37.42 (Rs 3,145).
Adani Power’s shares dropped more than 10% to $7.37 (Rs 619), but soon recovered to $8.04 (Rs 675)—down 2.9% from Friday’s closing.
Shares of the group’s other key listed firms—Adani Energy, Adani Green, Adani Total Gas and Adani Ports—were also hit on Monday, falling between 1-4%.
In a report published Saturday, New York-based short seller Hindenburg Research alleged Madhabi Puri Buch, the chair of the Securities and Exchange Board of India (SEBI), and her husband, Dhaval Buch, invested in offshore funds linked to Adani in Bermuda and Mauritius.
Adani’s brother Vinod Adani allegedly used these funds to purchase and trade “large positions in shares of the Adani Group,” the report alleged.
Get Forbes Breaking News Text Alerts: We’re launching text message alerts so you'll always know the biggest stories shaping the day’s headlines. Text “Alerts” to (201) 335-0739 or sign up here .
Forbes Valuation
According to our estimates , Gautam Adani’s net worth is $83.8 billion, down $1.5 billion due to Monday’s selloff. Despite this, he remains both India’s and Asia’s second richest person behind Mukesh Ambani.
Buch and her husband made the investments in 2015—two years before she joined SEBI, the report said, citing “whistleblower documents.” The Adani Group has dismissed Hindenburg’s allegations calling them “malicious, mischievous and manipulative selections of publicly available information to arrive at pre-determined conclusions for personal profiteering with wanton disregard for facts and the law.”
Key Background
Hindenburg disclosed a short position against the Adani Group’s listed firms in January 2023 and published a report accusing the conglomerate of engaging in “brazen stock manipulation and accounting fraud scheme over the course of decades.” The report caused Adani’s fortune to tank sharply, dropping from a high of $126 billion in January 2023 to less than $50 billion later in the year. The company vehemently denied these allegations and its billionaire founder labeled them a “malicious” attack on his company and India’s economic growth. The controversy spiraled into a political issue in India due to Adani’s decades-long close relationship with Indian Prime Minister Narendra Modi. Earlier this year, the Supreme Court rejected a request for an independent probe into Hindenburg’s allegations and the short seller was hit with a show cause notice by SEBI for violating Indian securities laws.
Chief Critic
Buch responded to Hindenburg’s allegations in a statement on Sunday: “The investment in the fund referred to in the Hindenburg report was made in 2015” when the Buchs were “both private citizens living in Singapore and almost 2 years before Madhabi joined SEBI.” The two consulting companies set up by Buch when she was in Singapore “became immediately dormant on her appointment with SEBI” and were “explicitly part of her disclosures.” The Buchs criticized Hindenburg, claiming the short seller had been served a “show cause notice for a variety of violations in India” and in response “they have chosen to attack the credibility of the SEBI and attempt character assassination of the SEBI Chairperson.” In response, Hindenburg tweeted : “Buch’s response now publicly confirms her investment in an obscure Bermuda/Mauritius fund structure, alongside money allegedly siphoned by Vinod Adani. She also confirmed the fund was run by a childhood friend of her husband, who at the time was an Adani director.”
Crucial Quote
India’s opposition parties criticized the government and have called for Buch’s resignation. Rahul Gandhi, the leader of the opposition in Parliament, responded to Hindenburg’s report: “This is an explosive allegation because it alleges the umpire herself is compromised. The savings of millions of Indians…are at risk, it is therefore imperative that this matter is investigated… If investors lose their hard-earned money, who will be held accountable—PM Modi, the SEBI Chairperson, or Gautam Adani?”
Further Reading
Indian Billionaire Adani Attacks 'Malicious' Hindenburg Report—And Invokes Nationalism Again (Forbes)
Adani Group Shares Slide After Hindenburg Alleges ‘Largest Con In Corporate History’ (Forbes)

- Editorial Standards
- Reprints & Permissions
Join The Conversation
One Community. Many Voices. Create a free account to share your thoughts.
Forbes Community Guidelines
Our community is about connecting people through open and thoughtful conversations. We want our readers to share their views and exchange ideas and facts in a safe space.
In order to do so, please follow the posting rules in our site's Terms of Service. We've summarized some of those key rules below. Simply put, keep it civil.
Your post will be rejected if we notice that it seems to contain:
- False or intentionally out-of-context or misleading information
- Insults, profanity, incoherent, obscene or inflammatory language or threats of any kind
- Attacks on the identity of other commenters or the article's author
- Content that otherwise violates our site's terms.
User accounts will be blocked if we notice or believe that users are engaged in:
- Continuous attempts to re-post comments that have been previously moderated/rejected
- Racist, sexist, homophobic or other discriminatory comments
- Attempts or tactics that put the site security at risk
- Actions that otherwise violate our site's terms.
So, how can you be a power user?
- Stay on topic and share your insights
- Feel free to be clear and thoughtful to get your point across
- ‘Like’ or ‘Dislike’ to show your point of view.
- Protect your community.
- Use the report tool to alert us when someone breaks the rules.
Thanks for reading our community guidelines. Please read the full list of posting rules found in our site's Terms of Service.
Recommended
Indian billionaire gautam adani loses another $2.4b after fresh hindenburg allegations.
Adani Group, the Indian conglomerate rocked by a Hindenburg Research report last year , faced another heavy share selloff on Monday after the shortseller accused the head of India’s market regulator of having links to offshore funds also used by the group.
About $2.4 billion, or 1%, had been wiped off the market value of Adani companies by the end of the trading day, although that was a substantial recovery from earlier losses of more than $13 billion.
The battle between Hindenburg Research and the Gautam Adani’s Adani Group began 18 months ago when the US shortseller alleged Adani improperly used tax havens, accusations the group denied again on Sunday, saying its overseas holding structure was fully transparent.

Citing whistleblower documents, Hindenburg said on Saturday that Madhabi Puri Buch, chair of the Securities and Exchange Board of India (SEBI) since 2022, has a conflict of interest in the Adani matter due to previous investments.
Buch said the report’s allegations were baseless and in a separate statement the regulator said allegations made by Hindenburg Research against the Adani Group have been duly investigated.
Shares in the group’s flagship firm Adani Enterprises closed out Monday 1.1% lower, while Adani Ports, Adani Total Gas, Adani Power, Adani Wilmar and Adani Energy Solutions were down between 0.6% and 4.2%. Only Adani Green bucked the trend, closing 1% higher.
“The allegations are coming for the second time. Lot of investigations have happened over the last year and a half. This is a temporary, knee-jerk reaction. Things will get back to normalcy,” said Sunny Agrawal, head of fundamental equity research at SBICAPS Securities.
Investments from Abu Dhabi-based International Holding and US boutique investment firm GQG Partners have helped restore some investor confidence since Hindenburg’s first report in January 2023, with Adani Group’s share value losses narrowing to about $32.5 billion from $150 billion in the immediate aftermath.
Buch termed Hindenburg’s allegations an attempt at “character assassination” following the regulator’s enforcement action and “show cause” notice to the shortseller for violating Indian rules.

A show cause notice signals an intention to take disciplinary action if satisfactory explanations are not provided.
Adani Enterprises is looking to launch a $1 billion share sale by mid-September, having shelved a record $2.5 billion offer in the wake of Hindenburg’s first set of allegations.
Adani Energy raised $1 billion from US investors and sovereign wealth funds earlier this month.
“We will likely see a short to medium term sentiment impact on Adani stocks, especially as retail investors are pressurized by the allegations made against SEBI,” said Kranthi Bathini, Director, Equity Strategy, WealthMills Securities.

As the latest allegations gained political traction, ruling Bharatiya Janata Party lawmaker Ravi Shankar Prasad said: “Instead of giving a response to the SEBI show cause notice, Hindenburg has issued this report, which is a baseless attack.”
“The SEBI and the family (of Buch) have responded, we don’t have anything to add to that,” he told reporters.
However, opposition leader Rahul Gandhi said on X: “The integrity of SEBI, the securities regulator entrusted with safeguarding the wealth of small retail investors, has been gravely compromised by the allegations against its chairperson.”

Advertisement
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 04 August 2024
Accurate prediction of protein function using statistics-informed graph networks
- Yaan J. Jang ORCID: orcid.org/0000-0002-0228-8672 1 , 2 na1 ,
- Qi-Qi Qin 2 , 3 na1 ,
- Si-Yu Huang 2 , 4 , 5 ,
- Arun T. John Peter 6 ,
- Xue-Ming Ding 3 &
- Benoît Kornmann ORCID: orcid.org/0000-0002-6030-8555 1
Nature Communications volume 15 , Article number: 6601 ( 2024 ) Cite this article
10k Accesses
21 Altmetric
Metrics details
- Bioinformatics
- Computational models
- Machine learning
- Protein function predictions
- Protein sequencing
Understanding protein function is pivotal in comprehending the intricate mechanisms that underlie many crucial biological activities, with far-reaching implications in the fields of medicine, biotechnology, and drug development. However, more than 200 million proteins remain uncharacterized, and computational efforts heavily rely on protein structural information to predict annotations of varying quality. Here, we present a method that utilizes statistics-informed graph networks to predict protein functions solely from its sequence. Our method inherently characterizes evolutionary signatures, allowing for a quantitative assessment of the significance of residues that carry out specific functions. PhiGnet not only demonstrates superior performance compared to alternative approaches but also narrows the sequence-function gap, even in the absence of structural information. Our findings indicate that applying deep learning to evolutionary data can highlight functional sites at the residue level, providing valuable support for interpreting both existing properties and new functionalities of proteins in research and biomedicine.
Similar content being viewed by others
LM-GVP: an extensible sequence and structure informed deep learning framework for protein property prediction
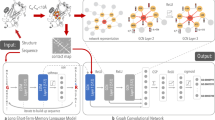
Structure-based protein function prediction using graph convolutional networks

Physicochemical graph neural network for learning protein–ligand interaction fingerprints from sequence data
Introduction.
Proteins bind to other molecules to facilitate nearly all essential biological activities. Consequently, understanding protein function is of paramount importance for comprehending health, disease, evolution, and the functioning of living organisms at the molecular level 1 , 2 , 3 . The primary sequence of a protein contains all the essential information required to fold up into a particular three-dimensional shape, thereby determining its activities within cells 4 , 5 . The evolutionary information in massive protein sequences that are gleaned from extensive genome sequencing efforts has significantly contributed to recent advances in protein structure prediction 6 , 7 , 8 , 9 . This evolutionary data, especially the couplings between pairwise residues, has also been utilized to characterize protein functional sites 10 , 11 . The evolutionary couplings have been utilized to pinpoint functional sites in proteins, capturing interactions between residues that contribute to specific functions 5 , 12 . Indeed, the analysis of evolutionary information has allowed the identification of allosteric mechanisms in proteins 13 , 14 , disease variants 15 , and metamorphism in proteins that undergo reversible switches between distinct folds, often accompanied by different functions 16 .
To date, more than 356 million proteins in the UniProt database 17 (6/2023) have been sequenced and the vast majority (~80%) of these have no known functional annotations (e.g., enzyme commission numbers and gene ontology terms). Classical methods for annotating protein functions have been constrained by the extensive sizes of sequences, and the majority of function annotations are assigned at the protein level rather than the residue level 18 , 19 . As an alternative to these classical methods, computational approaches have been utilized to assign function annotations to proteins 20 , 21 , 22 , 23 , 24 . Notably, deep learning methods have attained remarkable accuracy in predicting protein 3D structures, surpassing the capabilities of classical approaches such as ab initio methods and homology modeling. These methods involve millions of parameters and operate without making any assumptions about the relationship between input and output data samples (e.g., AlphaFold 8 and RoseTTAFold 9 ). Unlike the classical approaches, deep learning-based methods learn a large amount of parameters directly through the training of neural networks on extensive datasets. This enables them to generate accurate mappings from input data to expected outputs. Yet accurately assigning function annotations to proteins remains challenging, especially in comparison to experimental determinations. While there is abundant data available–whether from a single amino acid sequence, alignments of numerous homologous sequences, or protein structural information–to train deep learning-based methods, achieving accurate protein function prediction remains a persistent challenge 20 , 21 , 22 , 23 , 24 , 25 . Integrating physics-based knowledge from provided datasets, physics-informed deep learning methods have driven recent advances across diverse fields 26 . As a promising alternative to classical and pure deep learning techniques, they enhance the capacity of machine learning to construct interpretable methods for scientific problems. Despite decades of dedicated effort, assigning a function to a protein is more arduous than predicting its 3D structure 21 , 27 , 28 , 29 , 30 . The state-of-the-art approaches that utilize structural information have encountered less success in accurately assigning protein functions 21 . This is largely attributed to the scarcity of experimentally determined protein structures in comparison to the abundance of available sequences. Moreover, computationally predicted structures vary in their confidence scores and may not always be reliable for estimating protein function annotations, leading to variable accuracy in function annotation 21 , 30 . Furthermore, assessing the significance of residues using a scoring function that reliably measures their contributions to function remains challenging, as a quantitative characterization of residue roles is not yet fully comprehended.
To address these challenges, we hypothesized that the information encapsulated in coevolving residues can be leveraged to annotate functions at the residue level. Here, we devised a statistics-informed learning approach, termed PhiGnet, to facilitate the functional annotation of proteins and the identification of functional sites. Our method capitalizes on the knowledge derived from evolutionary data to drive two stacked graph convolutional networks. Empowered by the acquired knowledge and designed network architecture, the present method can accurately assign function annotations to proteins and, importantly, quantify the significance of each individual residue with respect to specific functions.
PhiGnet for protein function annotations
In this study, we developed the PhiGnet method using statistics-informed graph networks to annotate protein functions and to identify functional sites across species based on their sequences (Fig. 1 ). To assimilate knowledge from the evolutionary couplings (EVCs, relationships between pairwise residues at two co-variant sites) and the residue communities (RCs, hierarchical interactions among residues) 12 , we devised the method with a dual-channel architecture, adopting stacked graph convolutional networks (GCNs) (Fig. 1 a). This method specializes in assigning functional annotations, including Enzyme Commission (EC) numbers and Gene Ontology (GO) terms (biological process, BP, cellular component, CC, and molecular function, MF), to proteins. When provided with a protein sequence, we derive its embedding using the pre-trained ESM-1b model 31 . Subsequently, we input the embedding as graph nodes, accompanied by EVCs and RCs (graph edges), into the six graph convolutional layers of the dual stacked GCNs. These layers, working in conjunction with a block of two fully connected (FC) layers, meticulously process the information from the two GCNs, ultimately generating a tensor of probabilities for assessing the viability of assigning functional annotations to the protein. In addition, an activation score, derived using the gradient-weighted class activation maps (Grad-CAMs) approach 32 , is used to assess the significance of each individual residue in a specific function. The score allows PhiGnet to pinpoint functional sites at the level of individual residues (bottom, Fig. 1 c, see Methods).
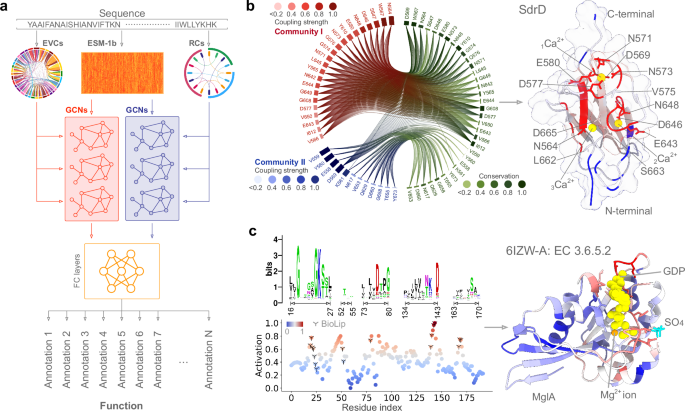
a PhiGnet predicts protein function from sequence alone. Given a sequence, PhiGnet learns the pre-embedding, EVCs, and RCs using stacked GCNs to infer protein function annotations. b RCs of the Serine-aspartate repeat-containing protein D (SdrD, PDB ID: 4JDZ). The two communities (community I and community II) with coupling strengths in bars are highlighted in red and blue. Each bar in either community I or II illustrates the strength of coupling that a residue has with others, while the conservation scores of these residues are depicted in the bars on the right. On the tertiary structure of SdrD (right), the residues within the community I (red) bind to the calcium ions (sphere in yellow) are shown in sticks, while the residues within the community II (blue) adopt cartoon in blue. c Function annotations of the MgIA protein at the residue level. The activation score (bottom) computed by PhiGnet is to measure the importance of each residue, where the higher the score is, the more likely it is to adopt a functional role in biological activity. Compared to functional sites in BioLip (marked with Y in black), the score indicates that co-evolved residues may be more important than those at conserved positions (top). The scores are mapped to color the MgIA 3D structure (PDB ID: 6IZW) from lower (blue) to higher (red), GDP is shown with sphere in yellow, SO 4 in stick in cyan, and Mg 2+ ion in a sphere with orange. Source data are provided as a Source Data file.
As an example, we computed RCs for the Serine-aspartate repeat-containing protein D (SdrD) that promotes bacterial survival in human blood by inhibiting innate immune-mediated bacterial killing 33 , 34 . Two RCs are mapped on a fully β sheet fold that binds to three Ca 2+ ions ( 1 Ca 2+ is enclosed in a loop, 2 Ca 2+ is more solvent exposed and closer to 3 Ca 2+ , which is coordinated by an asparagine (N564) and an aspartic acid (D665), Fig. 1 b). Within the community I, most residues (in red sticks) that are identified from EVCs bind to the three Ca 2+ ions, contributing together to stabilize the SdrD fold. This suggests that EVCs contain the essential information for deducing the functional role of residues, even when they are sparsely distributed across RCs. Empowered by EVCs and RCs, we implemented the present PhiGnet to assess the functional significance of residues. We carried out PhiGnet to calculate the activation scores for the functional sites of the mutual gliding-motility (MgIA) protein (annotated with EC 3.6.5.2) (Fig. 1 c). The resulting activation scores show that the residues with high scores ( ⩾ 0.5) are in agreement with or close to that of semi-manually curated BioLip database 35 . Moreover, these residues are located at the most conserved positions (top left, Fig. 1 c). Upon mapping these scores onto the 3D structure of MgIA, the activation scores highlight residues (red) that constitute a pocket that binds the guanosine di-nucleotide (GDP) and play a role in facilitating nucleotide exchange 36 . Together, this suggests that residues at functional sites are conserved through natural evolution, and that PhiGnet is capable of capturing such information, improving the method for predicting protein function at the residue level, even in the absence of structural data.
PhiGnet annotates protein functional sites
Many proteins perform their biological functions through essential residues that are sparsely distributed across different structural levels (e.g., primary, secondary, and tertiary) and are linked to functional sites (such as enzyme active sites, ligand-binding sites, or protein-protein interaction sites). Given the functional contributions of amino acids can significantly differ from one function to another, a key feature of PhiGnet is its ability to quantitatively estimate the importance of individual amino acids for a specific function, enabling us to identify residues that are pertinent to distinct biological activities.
Are the computational predictions as accurate as experimentally determined function annotations? To address this question, we carried out quantitative examinations of the contribution of each amino acid to a protein function using the activation score. We evaluated the predictive performance of PhiGnet and assessed the importance of residues (their contributions to protein function) in nine proteins: the c2-domain of cytosolic phospholipase A 2 α (cPLA 2 α ), Tyrosine-protein kinase BTK (TpK-BTK), Ribokinase, alpha-lactalbumin ( α LA), MCM1 transcriptional regular (MCM1-TR), the Fos-Jun heterodimer (FosJun), the thymidylate kinase (TmpK), Ecl18kI, and helicobacter pylori uridylate kinase (HPUK). These proteins vary in size from approximately 60 to 320 residues, harbor different folds, and perform diverse functions, including ligand binding, ion interaction, and DNA binding. We calculated the activation score for each residue in the nine proteins, comparing them to residues identified through either experimental or semi-manual annotations. Our method demonstrated promising accuracy (with an average ⩾ 75%) in predicting significant sites at the residue level, in a good agreement with actual ligand-/ion-/DNA-binding sites (Fig. 2 ). The activation score per residue, mapped onto their 3D structures, exhibits significant enrichment for functional relevance at the binding interfaces. PhiGnet accurately identifies functionally significant residues with high activation scores for the proteins (Fig. 2 , Supplementary Figs. S1 and S2) .
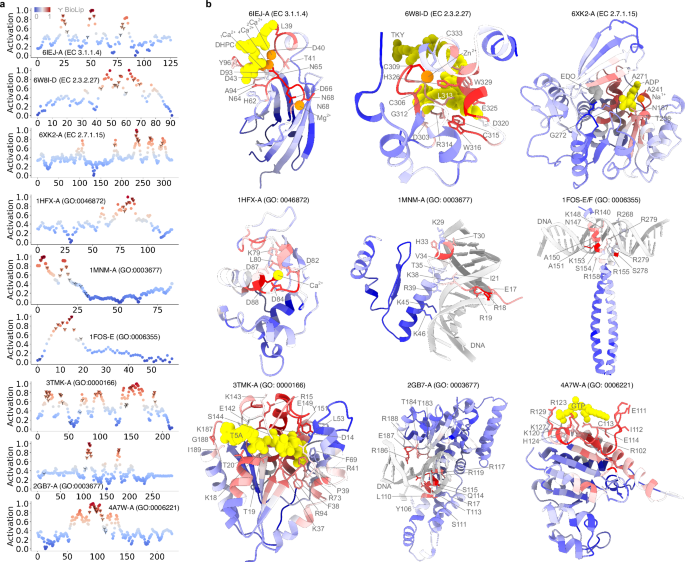
a The activation score of each residue is predicted using PhiGnet and compared to the biologically relevant ligand-protein binding sites from the BioLiP database. b The activation scores are mapped to the tertiary structures of nine proteins, including (left to right, top to bottom) the c2-domain of cytosolic phospholipase A 2 α (cPLA 2 α , PDB ID: 6IEJ) 37 , Tyrosine-protein kinase BTK (TpK-BTK, PDB ID: 6W8I), Ribokinase (PDB ID: 6XK2), alpha-lactalbumin ( α LA, PDB ID: 1HFX) 38 , MCM1 transcriptional regular (MCM1-TR, PDB ID: 1MNM) 60 , the Fos-Jun heterodimer (FosJun, PDB ID: 1FOS) 61 , the thymidylate kinase (TmpK, PDB ID: 3TMK) 62 , Ecl18kI (PDB ID: 2GB7) 39 , and helicobacter pylori uridylate kinase (HPUK, PDB ID: 4A7W) 63 . Source data are provided as a Source Data file.
Across the proteins cPLA 2 α , Ribokinase, α LA, TmpK, and Ecl18kI, PhiGnet predicted near-perfect functional sites compared to the experimental identifications. For instance, for cPLA 2 α , our method accurately identified residues, Asp40, Asp43, Asp93, Ala94 and Asn95, that bind to 1 Ca 2+ and residues, Asp40, Asp43, Asn65 and Thr41, that bind to 4 Ca 2+ , as well as a residue Asn65 supports 3 Ca 2+ for stabilizing fold 37 . Moreover, our method predicted a high score (0.6) for the residue Tyr96, which plays a crucial role in lipid headgroup recognition through cation- π interaction with the phosphatidylcholine trimethylammonium group 37 . We also applied PhiGnet to α LA, which contains a single, tightly bound calcium ion that is cradled in the EF-hand motif to stabilize the protein against denaturation 38 . In the α LA protein, the important motif is computationally characterized by a constellation of residues: Lys79, Asp82, Asp84, Asp87, and Asp88. In Ecl18kI, the major groove contacts the bases of the recognition sequence through the three consecutive residues Arg186, Glu187 and Arg188. Specifically, Arg186 and Arg188 form bidentate hydrogen bonds to the outer and inner guanines, respectively. The side chain oxygen atoms of Glu187 each accept one hydrogen bond from the two neighboring cytosines of the recognition sequence. Moreover, the sequence-specific minor groove contacts are exclusively mediated by Glu114 39 . To evaluate the importance of each residue in Ecl18kI, we computed the activation scores for each residue. These scores confirmed the agreement between the residues captured by PhiGnet and those identified through experimental data. For the proteins MCM1-TR and FosJun, our method captured residues with top activation scores that bind to DNAs, although not all of the residues at functional sites were characterized by high probabilities. Meanwhile, the activation scores failed to highlight function-relevant sites for a few residues. For instance, few residues with scores >0.5 were not located at the functional sites in Ribokinase, α LA, and HPUK. This discrepancy could be attributed to the noise present in EVCs. Together, the activation scores can indicate essential ligand-/ion-contacting residues, suggesting that learning from diverse levels of evolutionary knowledge can identify binding interfaces at the residue level. Such capability would be valuable in discerning interfaces both inter- and intra-proteins, even in the absence of structural information. Moreover, the predictions suggest that learning from evolutionary knowledge enables us to understand residues arranged in highly ordered patterns, relevant to diverse binding activities. On the other hand, biases originating from the evolutionary data could obscure the activation scores for accessing the functional significance of residues. Collectively, the activation scores can underscore essential ligand-/ion-contacting residues, indicating that learning from diverse levels of evolutionary knowledge can effectively identify binding interfaces at the residue level. Conversely, noise originating from the evolutionary data could influence the activation scores, potentially leading to biases in the identification of functional sites.
PhiGnet outperforms other state-of-the-art methods
To assess the predictive performance of PhiGnet, we implemented the method to infer function annotations (EC numbers and GO terms) for proteins in the two benchmark test sets (see Methods). We proceeded to compare our method against state-of-the-art methods, including alignment-based methods (BLAST 18 , FunFams 40 , and Pannzer 41 ), deep learning-based methods (DeepGO 25 , DeepFRI 21 , DeepGOWeb 42 , ProteInfer 43 , SPROF-GO 44 , ATGO+ 45 , and CLEAN 46 ). Two essential metrics, including the protein-centric F m a x -score and the area under the precision-recall curve (AUPR), were utilized for the comparisons. Our method demonstrated predictive capabilities for assigning function annotations to proteins across the two test sets. It achieved an average AUPR of 0.70 and 0.89, as well as F m a x scores of 0.80 and 0.88, for GO terms and EC numbers, respectively (Fig. 3 ). Moreover, it consistently maintained strong performance, with average AUPR scores of 0.64, 0.65, and 0.80, alongside corresponding F m a x values of 0.82, 0.75, and 0.81, for the three branches of GO terms – CC, BP, and MF (Fig. 3 d). Overall, PhiGnet significantly outperformed all supervised and unsupervised approaches across the benchmark datasets. For example, in the benchmark of EC numbers, we compared the predictions of various methods, including BLAST, FunFams, DeepGO, DeepFRI, Pannzer, ProteInfer, and CLEAN, against experimentally determined function annotations across the test proteins. Our method yielded F m a x score of 0.88 and AUPR of 0.89, surpassing the performance of other approaches (Fig. 3 a, b, Supplementary Fig. S3) . All the compared methods exhibited various performances, as illustrated in the precision-recall curves. DeepFRI, Pannzer, and ProteInfer achieved a similar F m a x score, approximately 0.68, outperforming BLAST and DeepGO. In terms of AUPR, FunFams, DeepFRI, and CLEAN yielded similar performances, which were better than those of ProteInfer and Pannzer. PhiGnet achieved F m a x of 0.88 and AUPR of 0.89, respectively, outperforming the CNN-based DeepGO (F m a x of 0.37 and AUPR of 0.21), structure-based DeepFRI (F m a x of 0.69 and AUPR of 0.70), and the contrastive learning-based CLEAN (F m a x of 0.76 and AUPR of 0.70) (Fig. 3 a, b, Supplementary Fig. S3) . These results suggest that PhiGnet has the ability to achieve accurate assignment of EC numbers to proteins. In the benchmark of GO terms, we compared our method against nine state-of-the-art methods, utilizing the same metrics to evaluate their performance. Across predictions of CC, BP, MF ontologies, PhiGnet achieved F m a x of 0.82, 0.75, 0.81 and AUPR of 0.64, 0.65, 0.80, respectively, which are significantly better than those of the compared methods. Notably, although ensemble-networks-based ProteInfer outperformed the remaining approaches over MF and BP ontologies, and the alignment-free SPROF-GO and structure-based DeepFRI excelled over CC ontology, PhiGnet’s performance remained superior (Fig. 3 d, e, Supplementary Figs. S4 – S7 , and Table S1) . Comparing predictive performances on the GO terms, we found that PhiGnet achieved first place in both accuracy and robustness, significantly better than the eight methods above and another prediction from a web server, DeepGOWeb (Fig. 3 d–f).
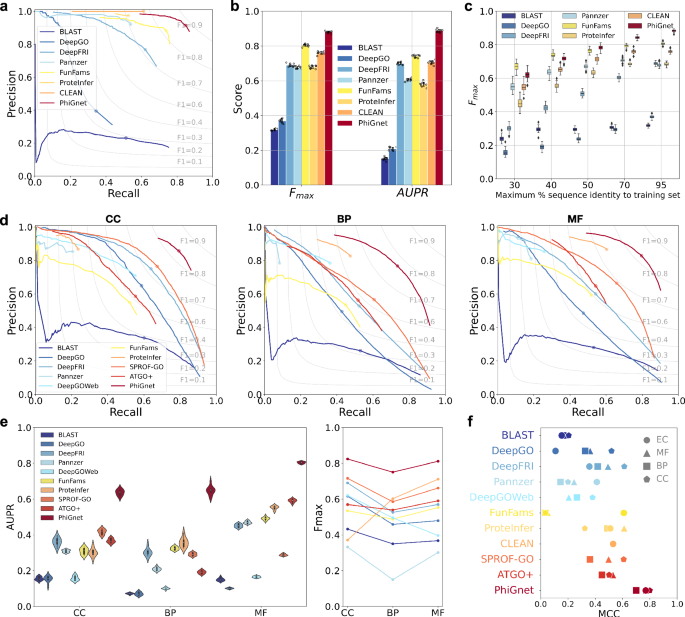
a Precision-recall curves illustrate the performance of different methods in predicting EC numbers for proteins. b Protein-centric F m a x scores and function-centric AUPR scores are computed across all test proteins to predict EC numbers, where the scores are presented as mean values with standard deviations of 10 bootstrap iterations. c Evaluation of robustness in predicting EC numbers as sequence identity increases, where the F m a x scores of each method at different sequence identities are depicted as boxplots of 50 bootstrap iterations, with the median values at the center and the interquartile range shown by the upper and lower edges of the boxes. d Precision-recall performance across GO terms in different ontologies. e Left, violin plots showing AUPR with the median values at the center of the distribution of 10 bootstrap iterations, and right, F m a x scores for the different methods in predicting CC, BP, and MF. f Computed Matthews correlation coefficient between predicted scores and ground-truth values for both EC numbers and GO terms using different methods. Source data are provided as a Source Data file.
Moreover, we demonstrated the robustness of PhiGnet for generalization to test proteins with varying thresholds of sequence identity compared to the proteins in the training set. At various maximum sequence identity levels (30%, 40%, 50%, 70%, and 95%), PhiGnet exhibited improved predictive performance as sequence identity increased (Fig. 3 c, Supplementary Fig. S5) . PhiGnet has been ranked among the top two robust methods for the test set of EC numbers, demonstrating consistently predictive performance with F m a x values of 0.61 and 0.72 at sequence identity levels of 30% and 40%, respectively. When compared to the domain-based method FunFams (F m a x of 0.67 and 0.74), PhiGnet slightly underperformed at sequence identity thresholds of 30% and 40%. However, PhiGnet achieved comparable or better performance when sequence identity exceeded 50%. Similarly, the performance of DeepFRI, FunFams, ProteInfer, and CLEAN also improved as sequence identity increased. Pannzer exhibited a similar trend when sequence identity was below 50%, but its performance remained nearly constant with a slight decrease in F m a x . In contrast, both BLAST and DeepGO showed slight improvements as the proteins in the test set increased sequence identity to those in the training set. The robustly predictive performance of PhiGnet has also been demonstrated by predicting the three branches of GO terms, maintaining high accuracy even at low sequence identity (Supplementary Fig. S5) . In predictions of both EC numbers and GO terms, we also calculated the Matthew’s correlation coefficient (MCC) between the predicted scores and ground truth to quantitatively compare the performance of various methods. PhiGnet achieved an average MCC of 0.76, which is higher than the average MCCs of the other ten state-of-the-art methods (Fig. 3 f).
PhiGnet driven by evolutionary signatures
The evolutionary data plays an important role in PhiGnet for predicting protein function annotations and identifying functional sites. First, we performed ablation experiments to test how EVCs/RCs contribute to PhiGnet. We trained PhiGnet using either EVCs or RCs alone and assessed its performance in terms of F m a x -score and AUPR over predictions of EC numbers/GO terms. To accomplish this, we chose a threshold (0.2) for both EVCs and RCs based on the similar performances in predicting EC numbers/GO terms (Supplementary Fig. S8) , aiming to mitigate potential noise arising from coevolution or weak couplings between pairwise residues. We first test whether the information in EVCs, which preserve evolutionary couplings at sites of co-variation, is sufficient to infer functional annotations. The second experiment tests the necessity of information in RCs that independently capture high-order couplings. Similarly, we built a model using RCs alone to computationally assign functional labels to proteins, and this model produced slightly better predictions (Supplementary Figs. S9 and S10) . The two experiments indicate that both models demonstrate the capability to accurately assign functional annotations to proteins. Moreover, PhiGnet, utilizing either EVCs or RCs, demonstrates a robust capacity to learn general sequence-function relationships, often better than or as good as other approaches, even test proteins exhibiting low sequence identity in presence of the training set (Fig. 3 c, Supplementary Figs. S9c and S10c) . Through precision and robustness comparisons, we have demonstrated that the evolutionary signatures (EVCs and RCs) constitute crucial attributes capable of enhancing deep learning-based methods for protein function annotations.
Secondly, we asked whether the residues, particularly within RCs that are often relevant to the specific function, can be quantified for functional sites. To address this, we further investigated the capability of PhiGnet to characterize meaningful features from the identified function-relevant residues within the residue communities. The activation scores were computed for the residues to underscore their contributions to the protein function. Notably, the predicted residues concurred with those at the functional sites identified through experimental determinations, better identifications than those in RCs (Fig. 4 ). In the human cytidine deaminase (hCDA) protein 47 , compared to residues within RCs that were identified as functionally relevant, PhiGnet quantitatively characterized their importance in the binding between hCDA and Zn 2+ /BRD through more accurate predictions of active sites: Cys65, Cys99, and Cys102, which coordinate with the zinc ion, as indicated by the activation scores (Fig. 4 a). In the Peroxide operon regulator (PerR), we also observed that PhiGnet narrowed down the number of residues located within RCs 48 and effectively distinguished non-Zn 2+ -binding residues from the binding ones, compared to RCs. Specifically, Cys96, Cys99 and Cys136, Cys139 exhibited much higher activation scores. These residues collectively coordinate the zinc ion, locking the three β -strands together to form the arrangement of the dimeric β -sheet, in contrast to the non-binding residues (Fig. 4 b). In light of these results, we conclude that the evolutionary information, particularly that contained in RCs, is sufficient to specify a protein’s function and to quantitatively characterize the residues at the functional sites. Moreover, the results argue that RCs contain evolutionary knowledge at a higher-ordered level than the information in EVCs at a lower-ordered level. Meanwhile, information contained in RCs plays an important role in enhancing PhiGnet’s ability to identify functionally relevant sites at the residue level.
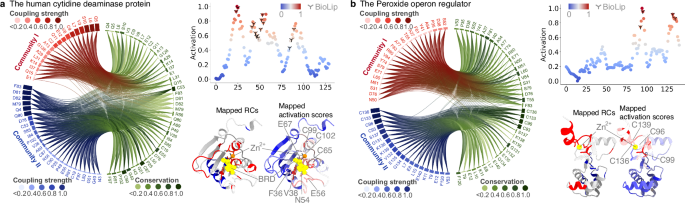
Mappings of RCs and activation scores of ( a ) the human cytidine deaminase protein (hCDA, PDB ID: 1MQ0-A, GO term 0008270), and ( b ) the Peroxide operon regulator (PDB ID: 2FE3-A, GO term 0046872). The residues within each RC are shown in the chord plotting with coupling strength and degree of conservation in bars. The activation scores (dotted lines) of each protein are compared to the BioLip identifications (marked with Y in black), and residues with high scores (in red) are also compared to those within RCs on their 3D structures. The 1-beta-ribofuranosyl-1,3-diazepinone (BRD) and Zn 2+ ions are shown with spheres in yellow (orange for the Zn 2+ ion in hCDA). Source data are provided as a Source Data file.
Test on CAFA3 targets
To assess whether the different performances of the methods under evaluation, and the superiority of PhiGnet were inherent to the algorithms or due to different training sets, we re-executed two alignment-based methods (BLAST and FunFams) and conducted retraining on four deep learning-based methods (DeepGO, ATGO+, SPROF-GO, and PhiGnet). Other methods were excluded primarily due to the unavailability of trainable source codes or because such method required unavailable structural information) against an identical dataset. We used the third Critical Assessment of Protein Function Annotation (CAFA3) dataset consisting of 66,841 proteins 49 . To address homology issues, proteins sharing over 30% sequence identity with the test proteins were excluded from the training dataset 45 . The remaining proteins were utilized to construct databases for BLAST and FunFams. 95% of them were randomly selected for training DeepGO, ATGO+, SPROF-GO, and PhiGnet, with the remaining 5% reserved for validation to fine-tune the methods’ parameters. Moreover, we conducted comparisons among the different methods using the CAFA3 test proteins either with less than 60% sequence identity to those in the training dataset or without redundancy removal (Supplementary Fig. S12) .
A comparison among the six different methods implemented on the CAFA3 dataset reveals that PhiGnet exhibits the best performance across both \({{{{\rm{F}}}}}_{\max }\) and AUPR metrics (Table 1 , Supplementary Fig. S12 ). PhiGnet achieved the highest \({{{{\rm{F}}}}}_{\max }\) scores across all three categories: BP (0.531), CC (0.584), and MF (0.606), indicating its superior capability in predicting functional annotations across diverse biological processes, cellular components, and molecular functions compared to methods such as BLAST, DeepGO, FunFams, and ATGO+. Furthermore, PhiGnet outperformed other methods with AUPR scores of 0.425 for BP, 0.590 for CC, and 0.571 for MF, demonstrating its effectiveness in accurately identifying true positive annotations while minimizing false positives across various functional categories. Although methods like BLAST, DeepGO, FunFams, and ATGO+ exhibited respectable performance in specific categories, none consistently achieved high scores across both \({{{{\rm{F}}}}}_{\max }\) and AUPR metrics as PhiGnet did. Overall, the comparison underscores PhiGnet as one of the state-of-the-art methods on the CAFA3 dataset, demonstrating that its increased performance is independent of the training dataset used.
Predicting functions of holdout and unannotated proteins
Can PhiGnet annotate uncharacterized proteins? We carried out our predictions for the independent hold-out set of 6229 proteins (Supplementary Fig. S13 ). We followed the same procedures to collect EVCs, RCs, and sequence embeddings for all the proteins. They were utilized to feed into the fine-tuned PhiGnet in order to compute a probability tensor for assigning functional annotations to the proteins. Among the collected proteins, our method’s overall performance was superior to that of state-of-the-art methods. Given that these proteins were independently collected, our computational predictions can be valuable in assigning functional annotations to new proteins (Supplementary Figs. S14 , S15 , and Table S2 ). For example, across the T. forsythia NanH (PDB ID: 7QXO) and human Sar1b (PDB ID: 8E0A), the activation scores successfully indicate the functional sites that bind to Oseltamivir and guanosine tetraphosphate (Supplementary Fig. S16) . Our analysis shows that PhiGnet’s high confidence prediction is in a good agreement with experimental annotations, suggesting that it would contribute to computational efforts for assigning function annotations to proteins with unknown labels. This applies even when dealing with experimental annotations of lower confidence scores, and can benefit experimental investigations of different biological activities. Moreover, by leveraging evolutionary information, PhiGnet provides function annotations as well as residue-level activation scores for over 2.5 million individual sequences within the UniProt database. The activation score assigned to each individual residue offers a quantitative measure of its significance in a specific activity, proving beneficial for screening experiments aimed at identifying functionally important sites.
It has been long appreciated that investigating evolutionary information across species can further our understanding of protein function and of the consequences of pathological mutations, even at the residue level. By leveraging deep learning methods on continuously expanding sequencing data, we can extract valuable knowledge to accurately annotate protein functions. This can greatly benefit both biological and clinical research, as well as facilitate drug discovery.
We have demonstrated that a statistics-informed learning method trained solely on evolutionary data achieves state-of-the-art performance in predicting protein function annotations at the residue level. The approach presented here requires no inputs other than the protein sequence and learns its characterized embedding using the statistics-informed graph convolutional networks. We show that EVCs and RCs have crucial effects on the predictions of protein function annotations and on the identifications of residues at functionally relevant sites. Our method produces high-accuracy annotations and identifies functional sites at the residue level. Therefore, this approach is well-suited for gaining a better understanding of the biological activities of unannotated or poorly studied proteins, as well as for quantitatively investigating the effects of disease-related variants.
When evaluating the performance of the methods presented (see Fig. 3 ), it becomes evident that PhiGnet outperforms its counterparts due to its distinctive amalgamation of two key factors. Firstly, it integrates insights derived from both evolutionary coupling analysis and spectrum analysis, resulting in a more comprehensive grasp of the intricate relationship between protein sequences and their functions. In contrast, other methods, such as FunFams and Pannzer, predominantly rely on homology-based approaches. Although homology-based methods have their merits, they might not capture the subtle nuances and intricate connections between proteins that are unveiled by the evolutionary coupling data. Conversely, while DeepFRI, DeepGO, SPROF-GO, and ATGO+ depend on structural data and homologous information, they may not harness the same depth of evolutionary data as PhiGnet. Moreover, the spectrum analysis applied to evolutionary data delves into the high-order patterns within protein sequences, which also contributes to PhiGnet’s superior performance. Secondly, although DeepFRI, DeepGO, SPROF-GO, ATGO+, and CLEAN are effective in leveraging pre-trained models for protein function prediction, PhiGnet distinguishes itself by enhancing the pre-trained model with evolutionary insights. This augmentation enables PhiGnet to offer a more holistic perspective on protein functions. By combining the ESM-1b model with evolutionary knowledge, PhiGnet achieves a deeper and more comprehensive understanding of the intricate relationship between protein sequences and their functions. This unique combination gives PhiGnet a competitive edge in accurately assigning EC numbers or GO terms to proteins, as it taps into a broader array of evolutionary features that many other methods do not fully explore.
In conclusion, the better performance of PhiGnet can be attributed to its utilization of the evolutionary data and high-order patterns of the data from protein sequences, allowing for a deeper and more accurate understanding of protein functions. PhiGnet leverages physically-inferred knowledge (EVCs and RCs) and performs significantly better predictions across both benchmark test sets of EC numbers and GO terms. This underscores PhiGnet’s capacity to effectively assimilate enriched evolutionary knowledge, where protein function has evolved and been encoded, to delineate the intricate relationship between protein sequences and their functions. Moreover, PhiGnet achieved higher accuracy in F m a x compared to the other approaches, even when dealing with proteins in the test set with low sequence identity to those in the training set. These comparisons lead us to conclude that PhiGnet demonstrates the capability for generalization in predicting protein function annotations across both EC numbers and GO terms.
The primary success of our approach lies in the utilization of statistics-informed graph convolutional neural networks to facilitate hierarchical learning over evolutionary data from massive sequence datasets. This approach surpasses existing supervised and unsupervised methods significantly and may be used to guide future biological and clinical experiments. We are aware that machine learning-based methods are highly dependent on the datasets that are used to tune their parameters. To mitigate bias arising from the datasets, it is important to curate proteins for training, maintain diversity in sequences, and evaluate the methods on various proteins to assess their generalization capabilities. Limitations of our method might include biases/noise arising in protein families with less diverse sequences. Incorporating (co-)evolutionary information into PhiGnet can impact the accurate identification of residue communities, particularly if the information is derived from a highly conserved protein family. While integrating physically extracted knowledge into our method yields a significant improvement compared to other approaches, there are still significant challenges in interpreting the learning mechanisms within PhiGnet. For instance, a protein might have more than one active or functionally relevant sites. The activation score does not allow to discern active site a given residue is part of.
We anticipate that evolutionary information will enable statistics-informed learning approaches to effectively characterize protein function at the residue level, including predicting disease variants, allosteric regulation, binding affinity, and specificity from sequence alone, as well as incorporating structural information for specific applications. The synergy between evolutionary data and machine learning will pave the way for accurately determining and engineering the biophysical properties of proteins, with implications spanning clinical decisions, industrial applications, and environmental biotechnology.
In the present study, we collected protein chains from the Protein Data Bank (PDB) 50 using the protocols 21 to construct datasets (until 10/2021). The collected protein chains were clustered at 95% sequence identity. From each cluster, we selected a representative protein possessing at least one annotated function. Two benchmark datasets were created, comprising 41,896 and 20,215 protein chains (with a maximum of 1024 residues each), annotated with GO terms and EC numbers, respectively. In the benchmark of EC numbers, we extracted unique annotations from the third-/fourth-level of the proteins, forming a total of six primary catalytic reaction classes: oxidoreductase, transferase, hydrolase, lyase, isomerase, and ligase. For the benchmark of GO terms, the three categories, BP, CC, and MF, are utilized to evaluate and compare the performance of various methods in this study. In the present study, we divided each dataset into three subsets, including training, validation, and test sets, with ratios of 8:1:1, respectively. The protein sequences in the test set (Supplementary Fig. S17) are of varying degrees (30%, 40%, 50%, 70%, and 95%) of sequence identity against that in the training set.
To create an independent hold-out set, we collected 13,584 proteins that are released after 1/2022 from the RCSB PDB database 50 (released between 1/2022 and 12/2022). Subsequently, we then searched these proteins against the SIFT database 51 (as of December 2022) to filter out proteins lacking experimentally determined functional annotations. As a result, we obtained 6229 proteins of less than 1024 residues as an independent hold-out test set. We implemented the trained PhiGnet to assign function annotations to these recently released proteins, and the predictions are evaluated against the annotations in the SIFT database.
Characterizing evolutionary signatures
To calculate evolutionary couplings, we collected an MSA for the target protein by searching its sequence against the UniClust30 database (up to February 2022) 52 using the hhblits tool 53 (version 3.3.0) with default parameters. Afterward, we performed trimming on each MSA using in-house scripts to eliminate sequences of low quality (for instance, sequences with over 80% gaps were removed). The distributions of MSA quality were obtained for both the training and test sets (Supplementary Fig. S18) . For each of the trimmed MSAs, we utilized our in-house scripts based on leri 12 to compute EVCs between pairwise residues. Subsequently, we derived RCs that capture functional signatures from these couplings. Both evolutionary couplings and residue communities were used as graph edges within PhiGnet in predicting protein functions. The computed EVCs may contain noise arising from the coevolution of residues across different sequences 54 . As a result, we implemented a normalization process on all computed EVCs, using a threshold of 0.2 to enhance their quality. Likewise, the scores within the RCs were also normalized to fall within the [0, 1] range and were subjected to filtering using a threshold of 0.2. These actions were informed by the experimental design’s focus on hyper-parameter optimization through grid search (Supplementary Fig. S8 ).
Learning information using the ESM-1b transformer
To allow evolutionary diversity of natural sequences, we leveraged the pre-trained model ESM-1b transformer 31 as physically embedded knowledge (across 250 million protein sequences) to improve the prediction ability of PhiGnet. The ESM-1b transformer is pre-trained on UniRef50 representative sequences and a specialized embedding of protein sequences to represent biological information at multiple levels, e.g., evolutionary homology. In this study, we derived the embedding of the provided protein sequence from the ESM-1b transformer’s output. This embedding was then integrated with EVCs and RCs to feed into PhiGnet. The integrated strategy offers insights into remote protein homology, leveraging informative relationships within the embedding representations of homologous proteins. This allows for generalization to previously unseen proteins in the training set.
We encoded each protein sequence using a sequence-level embedding from the ESM-1b model. Each amino acid is represented by a one-hot feature vector and embedded as an input representation for PhiGnet. The ESM-1b embedding captures the unique amino acid at each specific site along the sequence, enabling the stacked GCN layers to acquire higher-level features from either EVCs or RCs using distinct convolutional filters.
Statistics-informed graph networks
PhiGnet adopts dual channels consisting of stacked GCNs. In one channel, a stack of GCNs gathers information from the sequence embedding using evolutionarily coupled residues as graph nodes. In the other channel, the graph layers learn information about functionally significant residues using RCs as nodes. The PhiGnet architecture is composed of six GCN layers and two fully connected layers with dropout. Initially, a protein sequence of interest is used to compute EVCs, RCs, and the ESM-1b embedding information 31 . The first layer of each channel loads tensors of L × 1,280 from sequence embedding, and a tensor of EVCs/RCs is used as the adjacency matrix throughout all the three stacked graph layers (Fig. 1 a). In the two channels, EVCs are to describe the linkage between pairwise residues, while RCs are used to characterize hierarchical interactions for the other three stacked graph layers (Supplementary Fig. S19 ). They motivate PhiGnet to learn knowledge of residues that significantly contribute to protein function. The final fully connected layer incorporates a fixed-number SoftMax layer to compute the prediction probability for assigning function annotations to the protein.
In PhiGnet, we embed the given sequence of L amino acids using the ESM-1b transformer as a tensor T e s m ( T e s m ∈ R L × D , D is the dimension of the tensor). The sequence embedding is the input of the two channels in GCN to represent graphs at different levels, and we employ two adjacency matrices (EVCs and RCs) to describe the linkages between residues at two different levels. In each GCN layer of PhiGnet, we employed an undirected connected graph G = { V , E , A }, consisting of a set of nodes V with L residues, a set of edges E defined by the adjacency matrix A (a matrix of EVCs or RCs is used in the present study). If residue i is correlated with residue j as defined by the entry A ( i , j ) = 1; otherwise, there is no edge between residues i and j , A ( i , j ) = 0. The degree of the matrix A is denoted as a diagonal matrix D , where \({{{\bf{D}}}}(i,\, i)={\sum }_{j=1}^{n}{{{\bf{A}}}}(i,\, \, j)\) . Each GCN layer involves two phases of aggregation, where each node gathers and aggregates features of its neighbor nodes to update the local features, and combination, where the updated features are further merged to extract high-level abstraction through a local multilayer perceptron network. The layer-wise forward propagation of GCN is defined as follows,
where H ( k ) and W ( k ) are the representation of residues and weights of the k th layer, respectively, and σ ( ⋅ ) non-linear activation functions. In the present study, we implemented a normalized form over GCN and essentially arrive at the propagation rule 55 :
with \(\hat{{{{\bf{A}}}}}={{{\bf{A}}}}+{{{\bf{I}}}}\) , where I is an identity matrix and \(\hat{{{{\bf{D}}}}}\) is the diagonal node degree matrix of \(\hat{{{{\bf{A}}}}}\) .
There are three blocks of GCN layer that are used in each channel of PhiGnet, and the number of hidden units in each GCN layer is set to 512. Information extracted by different channels, using either EVCs or RCs, can promote PhiGnet to learn features at two levels (Supplementary Figs. S9 – S11) . The outputs of the GCNs are concatenated into a tensor of dimensions L × D , where L represents the number of nodes in the graphs. To consolidate the information across the L dimension, we apply a SumPooling layer, reducing L to 1 while preserving the other dimension. This aggregated tensor of size 1 × D is forwarded to the FC layers for predicting protein functions.
Hyper-parameter tuning and PhiGnet training
The present PhiGnet allows us to directly learn information from a sequence alone (without using any structural knowledge) to significantly explore functional sites at the residue level. To achieve an optimized model, we have to tune and choose values of the hyper-parameters in our method, e.g., thresholds for filtering EVCs/RCs (Supplementary Fig. S8 ). This tuning of parameters is crucial to guarantee both the stability and performance of PhiGnet.
With the pre-defined hyper-parameters, we implemented a cross-entropy loss function to balance the abilities of learning and generalization. The loss function is defined as follows,
where N is the number of data samples, and F is the number of function classes in EC numbers/GO terms. y i j is to label the ground truth to 1 if the i th sample is in the j th function class, otherwise, it is 0. Similarly, \({\hat{y}}_{ij}\) is a label for the prediction.
PhiGnet was trained with batch size of 64 for maximum 500 epochs using early-stopping criterion over the defined cross-entropy loss (Eq. ( 3 )). During training, we leveraged the Adam optimizer 56 with a learning rate of 2 × 10 −4 , β 1 = 0.9, β 2 = 0.999, ϵ = 1 × 10 −6 , and L 2 weight decay of 2 × 10 −5 . To avoid over-fitting, we implemented a dropout of 0.3 for the second fully connected layer. Accordingly, we achieved fine-trained models of PhiGnet that are leveraged to predict the probability of assigning EC numbers/GO terms to a given protein by learning from sequence embedding under constraints of evolutionary couplings and couplings intra residue communities.
Function annotations at the residue level
To quantitatively evaluate the importance of residues, we implemented the gradient-weighted class activation map method (that localizes the most important regions in images relevant for making correct classification decisions in computer vision) 32 for a specific function annotation to compute scores that are assigned to each residue in a protein. In the grad-CAM method, the gradient information of a given layer is used to compute localization map \({{{{\bf{M}}}}}^{c}\in {{\mathbb{R}}}^{u\times v}\) with width u and height v , and it is used to characterize the importance of every single element of the input for a specific class c . Given a feature map F k , the activation value \({{{{\mathcal{S}}}}}^{c}\) for scoring the class c is computed to measure the importance of neurons, \({\alpha }_{k}^{c}\) , as follows,
where ReLU( ⋅ ) is a non-linear activation function, holding a positive effect for function class c , and L is the number of elements in the input.
In the present method, we evaluated the importance of the i th amino acid in the feature map F k obtained from the layer concatenated from the two channels in PhiGnet, and the gradient \(\frac{\partial {{{{\bf{Y}}}}}^{c}}{\partial {{{{\bf{F}}}}}_{i}^{k}}\) is calculated by the derivative of the function annotation c with predicted score Y c , with respect to the feature map \({{{{\bf{F}}}}}_{i}^{k}\) in sequence of length L .
Comparison with existing approaches
In the present study, we compared our method to eight methods, including BLAST 18 , FunFams 40 , DeepGO 25 , DeepFRI 21 , ProteInfer 43 ATGO 45 , SPROF-GO 44 , and CLEAN 46 in details. Moreover, our method was compared to predictions collected from two web-servers, DeepGOWeb 42 and Pannzer 41 , over predictions of either GO terms in different ontologies or EC numbers using the collected data sets.
BLAST is a sequence searching tool based on the local sequence alignment algorithm 18 . Implementing BLAST, we transferred function annotations to proteins within the test set from all the annotated sequences in the training dataset following the same procedure as presented in refs. 20 , 21 . The probability assigning annotation(s) to each protein was computed by sequence identity in percentage between the sequences in the test and training sets. More specifically, if a protein in the test set hits against proteins in the training set with a maximum sequence identity of 75%, it was assigned function annotation(s) by transferring all the annotations from training proteins with a score of 0.75. In practice, we filtered out sequences from the training set using default parameters to keep within limits of assigning annotation(s) from homologous sequences 21 .
FunFams is a domain-based approach that leverages CATH super-families to transfer function annotation from a protein to another 40 . Given a protein, its sequence is searched against the CATH using the HMMER tool 57 , and its function annotation (EC numbers and GO terms) is copied from the FunFams with the highest HMM score. We obtained EC numbers and GO terms for the test proteins in this study by following the procedure present at https://github.com/UCLOrengoGroup/cath-tools-genomescan . More specifically, each protein is assigned a score (measuring either GO terms or EC numbers) that is computed from the frequency of proteins from the sequence alignment collected by FunFams from the CATH database.
DeepGO is a supervised deep learning method using convolutional neural networks (CNN) to predict GO terms initially 25 . DeepGO learns features from both protein sequences and a cross-species protein-protein interaction network using a CNN layer with 32 filters. In DeepGO, each protein sequence is encoded as a one-hot embedding and fed into the CNN model to compute sequence representation, combined with the embedding of protein-protein network. With a fully-connected layer of a sigmoid activation function, DeepGO generates a probability as confidence to assign a function annotation the query sequence. For fair comparison, we locally adopted DeepGO with default settings to predict both EC numbers and GO terms for the test set of proteins.
DeepFRI was constructed based on an architecture of graph convolutional networks to learn both protein sequence using a pre-trained LSTM model and its structural information 21 . DeepFRI leverages the pre-trained LSTM model to extract the feature of sequence, and such feature is learned by the graph convolutional networks using residue contacts that are derived from protein tertiary structure as representations for connections of residues, e.g., the i th and j th residues are contacted if the distance between the two C α atoms of the residues is less than a threshold of 10 Å; otherwise, they are not contacted. We locally implemented DeepFRI with its default configurations and collected the protein structures for the test set from the RCSB PDB database 50 . The residue contacts within each protein were computed under the threshold from its structure and used as structural information for DeepFRI to predict EC numbers/GO terms.
DeepGOWeb is developed based on DeepGOPlus 58 , an extended variant of the DeepGO method, and it utilizes many convolutional filters of different kernel sizes to learn protein sequence representations. As an improved method, it further embeds homology-based predictions from DIAMOND 59 to improve predictive accuracy. We collected the DeepGOWeb predictions over our test set of proteins from its webs-server with default parameters. We submitted our test protein sequences to the DeepGOWeb web-server and collected the predictions over the test sequences to compute both protein-centric F m a x score and term-centric AUPR for comparison.
Pannzer is a weighted K-nearest neighbor predictor for assigning function annotations to proteins 41 . Pannzer searches a query sequence against the UniProt database to collect the sequence neighborhood, and the annotations are transferred to the query protein from its homologous neighbors. We collect the Pannzer predictions of EC numbers and GO terms on our test set using its web-server.
ProteInfer is a method based on a single convolutional neural network scan for all known domains in parallel 43 . Proteinfer has 1100 filters to learn the mapping between protein sequences and functional annotations. The method was trained on the well-curated portion of Swiss-Prot data. The finely-tuned ProteInfer maps an amino acid sequence through five residual convolutional layers to create embeddings. These embeddings are then extracted using a fully connected layer featuring an element-wise sigmoid activation function, which facilitates the prediction of per-label probabilities.
SPROF-GO is a sequence-based alignment-free protein function predictor that embeds protein sequences using a pre-trained protein language model 44 . The sequence embedding is acquired through two parallel multi-layer perceptron networks, each designed for different latent representations. Additionally, another multi-layer perceptron is to map these representations to protein function label(s) (GO terms). The final predicted annotations are derived from the network model’s predictions and homology information with the training dataset, established using DIAMOND 59 .
ATGO adopts a triplet neural-network architecture using embeddings from the pre-trained ESM-1b model 31 to predict protein annotations (GO terms) 45 . In ATGO, the embeddings are generated from the last three layers and fused by a fully connected neural network. The triplet neural-network maps the fused representation to predict the confidence scores of protein GO terms. The ATGO+ method is a combination of the ATGO method and a sequence homology-based method, resulting in superior performance compared to ATGO.
CLEAN has been developed based on the contrastive learning for predictive assignments of EC numbers to enzymes 46 . The CLEAN method learns embedded representations of enzymes, in which proteins of the same EC numbers are close to each in Euclidean distances; otherwise, they are far from each other. The positive and negative samples are defined by the distances to the anchor sequence. Positive samples are closer to the anchor sequence, while negative samples are farther away from the anchor sequence. All sequences are embedded using the pre-trained ESM-1b model 31 and are then fed into a supervised contrastive learning neural network. Both the maximum separation and P value methods are employed to prioritize confident predictions of EC numbers in the final inferred results.
Performance metrics
We evaluate the different methods using two metrics: protein-centric maximum F-score (F m a x ) that measures the precision of labeling EC numbers/GO terms to a protein and term-centric area under precision-recall (AUPR) curve that measures the precision of labeling proteins to different EC numbers/GO terms. The F-score is the harmonic mean of the precision p ( t ) and recall r ( t ), while F m a x represents the maximum F-score achieved. F m a x and AUPR were defined as follows,
where p and r are precision that measures the predictive accuracy and recall that is to measure successfully retrieved information, respectively.
Statistics and reproducibility
No statistical method was used to predetermine sample size.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All relevant data supporting the key findings of this study are available within the article and its Supplementary Information files. All crystal structures of proteins used in this study are available at Protein Data Bank ( https://www.rcsb.org ) under accession codes: 4JDZ [ https://doi.org/10.2210/pdb4JDZ/pdb ], 6IZW [ https://doi.org/10.2210/pdb6IZW/pdb ], 6IEJ [ https://doi.org/10.2210/pdb6IEJ/pdb ], 6W8I [ https://doi.org/10.2210/pdb6W8I/pdb ], 6XK2 [ https://doi.org/10.2210/pdb6XK2/pdb ], 1HFX [ https://doi.org/10.2210/pdb1HFX/pdb ], 1MNM [ https://doi.org/10.2210/pdb1MNM/pdb ], 1FOS [ https://doi.org/10.2210/pdb1FOS/pdb ], 3TMK [ https://doi.org/10.2210/pdb3TMK/pdb ], 2GB7 [ https://doi.org/10.2210/pdb2GB7/pdb ], 4A7W [ https://doi.org/10.2210/pdb4A7W/pdb ], 1MQ0 [ https://doi.org/10.2210/pdb1MQ0/pdb ], 2FE3 [ https://doi.org/10.2210/pdb2FE3/pdb ], 7QXO [ https://doi.org/10.2210/pdb7QXO/pdb ], and 8E0A [ https://doi.org/10.2210/pdb8E0A/pdb ]. The data is available for download at https://doi.org/10.5281/zenodo.12496869 . Source data are provided with this paper.
Code availability
The PhiGnet Python code and pre-trained model are available at: https://doi.org/10.5281/zenodo.12496869 .
Repecka, D. et al. Expanding functional protein sequence spaces using generative adversarial networks. Nat. Mach. Intell. 3 , 324–333 (2021).
Article Google Scholar
Ferruz, N. et al. From sequence to function through structure: deep learning for protein design. Comput. Struct. Biotechnol. J. 21 , 238–250 (2022).
Boike, L., Henning, N. J. & Nomura, D. K. Advances in covalent drug discovery. Nat. Rev. Drug Discov. 21 , 881–898 (2022).
Article PubMed PubMed Central CAS Google Scholar
Anfinsen, C. B. The formation and stabilization of protein structure. Biochem. J. 128 , 737 (1972).
Socolich, M. et al. Evolutionary information for specifying a protein fold. Nature 437 , 512–518 (2005).
Article ADS PubMed CAS Google Scholar
Hopf, T. A. et al. Three-dimensional structures of membrane proteins from genomic sequencing. Cell 149 , 1607–1621 (2012).
Kuhlman, B. & Bradley, P. Advances in protein structure prediction and design. Nat. Rev. Mol. Cell Biol. 20 , 681–697 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596 , 583–589 (2021).
Article ADS PubMed PubMed Central CAS Google Scholar
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373 , 871–876 (2021).
Humphreys, I. R. et al. Computed structures of core eukaryotic protein complexes. Science 374 , eabm4805 (2021).
Wang, J. et al. Scaffolding protein functional sites using deep learning. Science 377 , 387–394 (2022).
Cheung, N. J., Peter, A. T. J. & Kornmann, B. Leri: a web-server for identifying protein functional networks from evolutionary couplings. Comput. Struct. Biotechnol. J. 19 , 3556–3563 (2021).
Changeux, J.-P. & Edelstein, S. J. Allosteric mechanisms of signal transduction. Science 308 , 1424–1428 (2005).
Faure, A. J. et al. Mapping the energetic and allosteric landscapes of protein binding domains. Nature 604 , 175–183 (2022).
Frazer, J. et al. Disease variant prediction with deep generative models of evolutionary data. Nature 599 , 91–95 (2021).
Dishman, A. F. et al. Evolution of fold switching in a metamorphic protein. Science 371 , 86–90 (2021).
Bateman, A. et al. UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res. 51 , D523–D531 (2023).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25 , 3389–3402 (1997).
Scheibenreif, L., Littmann, M., Orengo, C. & Rost, B. FunFam protein families improve residue level molecular function prediction. BMC Bioinforma. 20 , 1–9 (2019).
Article CAS Google Scholar
Radivojac, P. et al. A large-scale evaluation of computational protein function prediction. Nat. Methods 10 , 221–227 (2013).
Gligorijević, V. et al. Structure-based protein function prediction using graph convolutional networks. Nat. Commun. 12 , 1–14 (2021).
Gelman, S., Fahlberg, S. A., Heinzelman, P., Romero, P. A. & Gitter, A. Neural networks to learn protein sequence–function relationships from deep mutational scanning data. Proc. Natl. Acad. Sci. USA 118 , e2104878118 (2021).
Bileschi, M. L. et al. Using deep learning to annotate the protein universe. Nat. Biotechnol. 40 , 932–937 (2022).
Article PubMed CAS Google Scholar
Unsal, S. et al. Learning functional properties of proteins with language models. Nat. Mach. Intell. 4 , 227–245 (2022).
Kulmanov, M., Khan, M. A. & Hoehndorf, R. DeepGO: predicting protein functions from sequence and interactions using a deep ontology-aware classifier. Bioinformatics 34 , 660–668 (2018).
Karniadakis, G. E. et al. Physics-informed machine learning. Nat. Rev. Phys. 3 , 422–440 (2021).
Pazos, F. & Sternberg, M. J. Automated prediction of protein function and detection of functional sites from structure. Proc. Natl. Acad. Sci. USA 101 , 14754–14759 (2004).
Gherardini, P. F. & Helmer-Citterich, M. Structure-based function prediction: approaches and applications. Brief. Funct. Genom. Proteom. 7 , 291–302 (2008).
Glazer, D. S., Radmer, R. J. & Altman, R. B. Improving structure-based function prediction using molecular dynamics. Structure 17 , 919–929 (2009).
Skolnick, J. & Brylinski, M. FINDSITE: a combined evolution/structure-based approach to protein function prediction. Brief. Bioinforma. 10 , 378–391 (2009).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl. Acad. Sci. USA 118 , e2016239118 (2021).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. In Proceedings of the IEEE International Conference on Computer Vision , 618–626 (2017).
Askarian, F. et al. Serine-aspartate repeat protein D increases Staphylococcus aureus virulence and survival in blood. Infect. Immun. 85 , e00559–16 (2017).
Milles, L. F., Unterauer, E. M., Nicolaus, T. & Gaub, H. E. Calcium stabilizes the strongest protein fold. Nat. Commun. 9 , 1–10 (2018).
Article ADS CAS Google Scholar
Yang, J., Roy, A. & Zhang, Y. BioLiP: a semi-manually curated database for biologically relevant ligand–protein interactions. Nucleic Acids Res. 41 , D1096–D1103 (2012).
Article PubMed PubMed Central Google Scholar
Baranwal, J. et al. Allosteric regulation of a prokaryotic small Ras-like GTPase contributes to cell polarity oscillations in bacterial motility. PLoS Biol. 17 , e3000459 (2019).
Hirano, Y. et al. Structural basis of phosphatidylcholine recognition by the C2–domain of cytosolic phospholipase A2 α . Elife 8 , e44760 (2019).
Pike, A. C., Brew, K. & Acharya, K. R. Crystal structures of guinea-pig, goat and bovine α -lactalbumin highlight the enhanced conformational flexibility of regions that are significant for its action in lactose synthase. Structure 4 , 691–703 (1996).
Bochtler, M. et al. Nucleotide flips determine the specificity of the Ecl18kI restriction endonuclease. EMBO J. 25 , 2219–2229 (2006).
Das, S. Functional classification of CATH superfamilies: a domain-based approach for protein function annotation. Bioinformatics 31 , 3460–3467 (2015).
Törönen, P. & Holm, L. PANNZER—a practical tool for protein function prediction. Protein Sci. 31 , 118–128 (2022).
Article PubMed Google Scholar
Kulmanov, M., Zhapa-Camacho, F. & Hoehndorf, R. DeepGOWeb: fast and accurate protein function prediction on the (Semantic) Web. Nucleic Acids Res. 49 , W140–W146 (2021).
Sanderson, T., Bileschi, M. L., Belanger, D. & Colwell, L. J. ProteInfer, deep neural networks for protein functional inference. Elife 12 , e80942 (2023).
Yuan, Q., Xie, J., Xie, J., Zhao, H. & Yang, Y. Fast and accurate protein function prediction from sequence through pretrained language model and homology-based label diffusion. Brief. Bioinforma. 24 , bbad117 (2023).
Zhu, Y.-H., Zhang, C., Yu, D.-J. & Zhang, Y. Integrating unsupervised language model with triplet neural networks for protein gene ontology prediction. PLOS Comput. Biol. 18 , e1010793 (2022).
Yu, T. et al. Enzyme function prediction using contrastive learning. Science 379 , 1358–1363 (2023).
Chung, S. J., Fromme, J. C. & Verdine, G. L. Structure of human cytidine deaminase bound to a potent inhibitor. J. Med. Chem. 48 , 658–660 (2005).
Traoré, D. A. et al. Crystal structure of the apo-PerR-Zn protein from Bacillus subtilis . Mol. Microbiol. 61 , 1211–1219 (2006).
Zhou, N. et al. The CAFA challenge reports improved protein function prediction and new functional annotations for hundreds of genes through experimental screens. Genome Biol. 20 , 1–23 (2019).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28 , 235–242 (2000).
Ng, P. C. & Henikoff, S. Predicting deleterious amino acid substitutions. Genome Res. 11 , 863–874 (2001).
Mirdita, M. et al. Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res. 45 , D170–D176 (2017).
Remmert, M., Biegert, A., Hauser, A. & Söding, J. HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 9 , 173–175 (2012).
Anishchenko, I., Ovchinnikov, S., Kamisetty, H. & Baker, D. Origins of coevolution between residues distant in protein 3D structures. Proc. Natl. Acad. Sci. USA 114 , 9122–9127 (2017).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. arXiv preprint arXiv:1609.02907 (2016).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7 , e1002195 (2011).
Article ADS MathSciNet PubMed PubMed Central CAS Google Scholar
Kulmanov, M. & Hoehndorf, R. DeepGOPlus: improved protein function prediction from sequence. Bioinformatics 36 , 422–429 (2020).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12 , 59–60 (2015).
Tan, S. & Richmond, T. J. Crystal structure of the yeast MAT α 2/MCM1/DNA ternary complex. Nature 391 , 660–666 (1998).
Glover, J. & Harrison, S. C. Crystal structure of the heterodimeric bZIP transcription factor c-Fos–c-Jun bound to DNA. Nature 373 , 257–261 (1995).
Lavie, A. et al. Crystal structure of yeast thymidylate kinase complexed with the bisubstrate inhibitor P 1-(5 ‘-Adenosyl) P 5-(5 ‘-Thymidyl) pentaphosphate (TP5A) at 2.0 Å resolution: Implications for catalysis and AZT activation. Biochemistry 37 , 3677–3686 (1998).
Chu, C.-H. et al. Structures of helicobacter pylori uridylate kinase: insight into release of the product UDP. Acta Crystallogr. D Biol. Crystallogr. 68 , 773–783 (2012).
Download references
Acknowledgements
This work was funded by Wellcome Trust (214291/Z/18/Z, to B.K.). We thank members of the Kornmann laboratory and AmoAi for many valuable discussions. Y.J.J. and Q.Q.Q. are supported by AmoAi.
Author information
These authors contributed equally: Yaan J. Jang, Qi-Qi Qin.
Authors and Affiliations
Department of Biochemistry, University of Oxford, Oxford, UK
Yaan J. Jang & Benoît Kornmann
AmoAi Technologies, Oxford, UK
Yaan J. Jang, Qi-Qi Qin & Si-Yu Huang
School of Optical-Electrical and Computer Engineering, University of Shanghai for Science and Technology, Shanghai, China
Qi-Qi Qin & Xue-Ming Ding
Oxford Martin School, University of Oxford, Oxford, UK
Si-Yu Huang
School of Systems Science, Beijing Normal University, Beijing, China
Institute of Biochemistry, ETH Zürich, Zürich, Switzerland
Arun T. John Peter
You can also search for this author in PubMed Google Scholar
Contributions
Y.J.J. led the research, conceived the end-to-end approach, designed experiments, financed the experiments, and wrote the manuscript. Q.Q.Q. collected the data, implemented the method, contributed with principal analysis and wrote the first draft. S.Y.H. conducted principal analysis over predictions. X.M.D. conducted data analysis. A.T.J.P. supported with principal analysis and wrote the manuscript. B.K. led the research, funding acquisition, contributed technical advice, and wrote the manuscript. All authors read the final manuscript.
Corresponding authors
Correspondence to Yaan J. Jang or Benoît Kornmann .
Ethics declarations
Competing interests.
Y.J.J. is a founder of AmoAi Technologies, UK. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interests.
Peer review
Peer review information.
Nature Communications thanks Guoxian Yu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information, peer review file, reporting summary, source data, source data, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/ .
Reprints and permissions
About this article
Cite this article.
Jang, Y.J., Qin, QQ., Huang, SY. et al. Accurate prediction of protein function using statistics-informed graph networks. Nat Commun 15 , 6601 (2024). https://doi.org/10.1038/s41467-024-50955-0
Download citation
Received : 17 May 2023
Accepted : 15 July 2024
Published : 04 August 2024
DOI : https://doi.org/10.1038/s41467-024-50955-0
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
By submitting a comment you agree to abide by our Terms and Community Guidelines . If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.
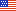

An official website of the United States government
Here’s how you know
Official websites use .gov A .gov website belongs to an official government organization in the United States.
Secure .gov websites use HTTPS A lock ( Lock A locked padlock ) or https:// means you’ve safely connected to the .gov website. Share sensitive information only on official, secure websites.

U.S. Department of Commerce
- Press Releases
Was this page helpful?
U.s. department of commerce announces preliminary terms with sk hynix to advance u.s. ai supply chain security, office of public affairs.
Biden-Harris Administration’s Bipartisan CHIPS and Science Act Attracts All Five Major Leading-Edge Logic and Memory Companies to Produce Chips on U.S. Soil
Today, the Biden-Harris Administration announced that the U.S. Department of Commerce and SK hynix have signed a non-binding preliminary memorandum of terms (PMT) to provide up to $450 million in proposed federal incentives under the CHIPS and Science Act to establish a high-bandwidth memory (HBM) advanced packaging fabrication and research and development (R&D) facility. President Biden signed the bipartisan CHIPS and Science Act to usher in a new era of semiconductor manufacturing in the United States, bringing with it a revitalized domestic supply chain, good-paying jobs, and investments in the industries of the future. The proposed CHIPS investment builds upon SK hynix’s investment of approximately $3.87 billion in West Lafayette, Indiana, to build a memory packaging plant for artificial intelligence (AI) products and an advanced packaging R&D facility, creating approximately 1,000 new jobs and filling a critical gap in the U.S. semiconductor supply chain.
“The Biden-Harris Administration’s CHIPS and Science Act is a once-in-a-generation opportunity to supercharge America’s global technology leadership and create quality jobs in the process. Today’s historic announcement with SK hynix would further solidify America’s AI hardware supply chain in a way no other country on earth can match, with every major player in advanced semiconductor manufacturing and packaging building or expanding on our shores,” said U.S. Secretary of Commerce Gina Raimondo. “Because of President Biden and Vice President Harris’ leadership, we are creating hundreds of new jobs in Indiana and ensuring the Hoosier state and Purdue University will play a crucial role in advancing America’s national security and supply chains.”
“President Biden and Vice President Harris are bringing the most advanced semiconductor manufacturing back to the United States,” said Arati Prabhakar, Assistant to the President for Science and Technology and Director of the White House Office of Science and Technology Policy. “Advanced packaging is more and more critical for AI and other leading-edge systems, but it requires extremely precise manufacturing processes. With this incentive from the CHIPS and Science Act, SK hynix will make a major contribution to the complex computing systems that our nation relies on. At the same time, we are making the R&D investments to win the future, too.”
The West Lafayette plant builds on SK Group’s previously announced multi-billion commitment to invest in American manufacturing, including EV batteries and biotechnology, which was announced during a meeting with President Biden in July 2022. Through the proposed CHIPS investment in SK hynix, the world’s leading producer of HBM, the Biden-Harris Administration would take a meaningful step in advancing the security of the U.S. AI supply chain. With this announcement, the United States will have preliminary agreements with all five of the world’s leading-edge logic, memory, and advanced packaging providers. No other economy in the world has more than two of these companies producing leading-edge chips on its shores.
SK hynix’s West Lafayette facility, located at the Purdue University Research Park, will be home to an advanced semiconductor packaging line that will mass-produce next generation HBM. These high-performance memory chips are crucial components of graphics processing units (GPUs) that train AI systems due to their increased processing power. This next generation chip would be mass-produced at the West Lafayette facility and will boast a more advanced performance than the company’s latest HBM, which processes up to 1.18 terabytes of data – the equivalent of 230 full HD movies – per second. Mass production at the facility is expected to begin in the second half of 2028.
“The Biden-Harris Administration is dedicated to inventing and commercializing semiconductor technology in the United States and to promoting domestic semiconductor manufacturing. With President Biden and Vice President Harris’ proposed investment in SK hynix, we could advance our commitment to accomplishing both,” said Under Secretary of Commerce for Standards and Technology and National Institute of Standards and Technology Director Laurie E. Locascio . “With proposed investments in companies like SK hynix, the United States has the opportunity to be the only country in the world where every company capable of producing leading-edge chips will have both a high-volume manufacturing presence and a significant research and development presence.”
As a result of this proposed investment, the Biden-Harris Administration would establish a research hub in Indiana because of SK hynix’s partnership with Purdue University, which hosts the largest facility of its kind at a U.S. university, while bringing next generation HBM and advanced packaging R&D to the United States. The next generation HBM that will be researched and developed, mass-produced, and packaged in this ecosystem with Purdue University will play an important role in the U.S. semiconductor ecosystem and advancing U.S. technological leadership.
“We deeply appreciate the U.S. Department of Commerce’s support and are excited to collaborate in seeing this transformational project fully realized,” said SK hynix CEO Kwak Noh-Jung. “We are moving forward with the construction of the Indiana production base, working with the State of Indiana, Purdue University and our U.S. business partners to ultimately supply leading-edge AI memory products from West Lafayette. We look forward to establishing a new hub for AI technology, creating skilled jobs for Indiana and helping build a more robust, resilient supply chain for the global semiconductor industry.”
SK hynix will collaborate with Purdue University on plans for future R&D projects, which include working on advanced packaging and heterogeneous integration with Purdue’s Birck Nanotechnology Center and other research institutes and industry partners. SK hynix plans to collaborate on projects for memory-centric solutions and architecture for generative AI – specifically memory design and in/near memory computing. As part of its workforce development efforts, SK hynix plans to work with Purdue University and Ivy Tech Community College to develop training programs and interdisciplinary degree curricula that will cultivate a high-tech workforce and build a reliable pipeline of new talent. Additionally, SK hynix plans to support the work of the Purdue Research Foundation and other local non-profits and charities by building partnerships that provide community development, growth opportunities, and leadership training.
The company has indicated that it plans to claim the Department of the Treasury’s Investment Tax Credit, which is expected to be up to 25% of qualified capital expenditures. In addition to the proposed direct funding of up to $450 million, the CHIPS Program Office would make up to $500 million of proposed loans – which is part of the $75 billion in loan authority provided by the CHIPS and Science Act – available to SK hynix under the non-binding PMT.
As explained in its first Notice of Funding Opportunity , the Department may offer applicants a PMT on a non-binding basis after satisfactory completion of the merit review of a full application. The PMT outlines key terms for a potential CHIPS incentives award, including the amount and form of the award. The award amounts are subject to due diligence and negotiation of award documents and are conditional on the achievement of certain milestones. After the PMT is signed, the Department begins a comprehensive due diligence process on the proposed projects and continues negotiating or refining certain terms with the applicant. The terms contained in any final award documents may differ from the terms of the PMT being announced today.
About CHIPS for America
Nearly two years after the passage of CHIPS and Science Act, the Biden-Harris Administration is moving full speed ahead in order to help protect our economic and national security and restore American leadership in an industry that we started decades ago. By allocating over $30 billion in proposed funding to build factories domestically and proposing to invest billions more in research and innovation, we are unlocking over $300 billion in public and private investment and creating more than 100,000 jobs, including tens of thousands of good-paying jobs that don't require a college degree. Our efforts are a meaningful step towards ensuring that the United States produces more of the world’s most advanced technologies – from AI to defense systems and everyday items like cars and medical devices. With a focus on expanding capacity, enhancing capabilities, maintaining competitiveness, and driving commercialization, CHIPS for America is working towards driving our future, securing our supply chains, and cementing America’s place at the forefront of technology.
CHIPS for America is part of President Biden’s economic plan to invest in America, stimulate private sector investment, create good-paying jobs, make more in the United States, and revitalize communities left behind. CHIPS for America includes the CHIPS Program Office, responsible for manufacturing incentives, and the CHIPS Research and Development Office, responsible for R&D programs, that both sit within the National Institute of Standards and Technology (NIST) at the Department of Commerce. Visit https://www.chips.gov to learn more.

IMAGES
COMMENTS
A research report is a collection of contextual data, gathered through organized research, that provides new insights into a particular challenge (which, for this article, is business-related). Research reports are a time-tested method for distilling large amounts of data into a narrow band of focus. Their effectiveness often hinges on whether ...
A research report is a well-crafted document that outlines the processes, data, and findings of a systematic investigation. It is an important document that serves as a first-hand account of the research process, and it is typically considered an objective and accurate source of information.
Thesis is a type of research report. A thesis is a long-form research document that presents the findings and conclusions of an original research study conducted by a student as part of a graduate or postgraduate program. It is typically written by a student pursuing a higher degree, such as a Master's or Doctoral degree, although it can also ...
An outline of the research questions and hypotheses; the assumptions or propositions that your research will test. Literature Review. Not all research reports have a separate literature review section. In shorter research reports, the review is usually part of the Introduction. A literature review is a critical survey of recent relevant ...
Research reports are recorded data prepared by researchers or statisticians after analyzing the information gathered by conducting organized research, typically in the form of surveys or qualitative methods. A research report is a reliable source to recount details about a conducted research. It is most often considered to be a true testimony ...
There are three main forms of reports: factual, instructional and persuasive; each has a different purpose and will require different arguments and evidence to achieve that purpose. It will help you write good reports if you know what you are trying to achieve before you start your report. Factual. Instructional. Persuasive.
THE RESEARCH REPORT. This chapter gives attention to two primary topics, both of which present information about research reports. The first part deals with the many valuable things that can be found in research reports beyond the obvious—the results. In the second part we discuss what a research report is and what it is not.
What are the implications of the findings? The research report contains four main areas: Introduction - What is the issue? What is known? What is not known? What are you trying to find out? This sections ends with the purpose and specific aims of the study. Methods - The recipe for the study. If someone wanted to perform the same study ...
This review is divided into sections for easy reference. There are five MAJOR parts of a Research Report: 1. Introduction 2. Review of Literature 3. Methods 4. Results 5. Discussion. As a general guide, the Introduction, Review of Literature, and Methods should be about 1/3 of your paper, Discussion 1/3, then Results 1/3.
A research report is one big argument about how and why you came up with your conclusions. To make it a convincing argument, a typical guiding structure has developed. In the different chapters, there are distinct issues that need to be addressed to explain to the reader why your conclusions are valid. The governing principle for writing the ...
Use the section headings (outlined above) to assist with your rough plan. Write a thesis statement that clarifies the overall purpose of your report. Jot down anything you already know about the topic in the relevant sections. 3 Do the Research. Steps 1 and 2 will guide your research for this report.
When reporting the methods used in a sample -based study, the usual convention is to. discuss the following topics in the order shown: Chapter 13 Writing a Research Report 8. • Sample (number in ...
A research report is a document that conveys the outcomes of a study or investigation. Its purpose is to communicate the research's findings, conclusions, and implications to a particular audience. This report aims to offer a comprehensive and unbiased overview of the research process, methodology, and results.
A research report is an end product of research. As earlier said that report writing provides useful information in arriving at rational decisions that may reform the business and society. The findings, conclusions, suggestions and recommendations are useful to academicians, scholars and policymakers.
A research report is a well-crafted document that outlines the processes, data, and findings of a systematic investigation. ... The framework furthermore serves at least the following functions ...
Table of contents. Step 1: Introduce your topic. Step 2: Describe the background. Step 3: Establish your research problem. Step 4: Specify your objective (s) Step 5: Map out your paper. Research paper introduction examples. Frequently asked questions about the research paper introduction.
Research Results. Research results refer to the findings and conclusions derived from a systematic investigation or study conducted to answer a specific question or hypothesis. These results are typically presented in a written report or paper and can include various forms of data such as numerical data, qualitative data, statistics, charts, graphs, and visual aids.
Comprehensive reports with in-depth analysis and information. 100-page research report on the effects of a new drug on a medical condition. Analytical. Focus on data analysis and provide insights or recommendations. Market research report analyzing consumer behavior trends and recommending marketing strategies.
1 Choose a topic based on the assignment. Before you start writing, you need to pick the topic of your report. Often, the topic is assigned for you, as with most business reports, or predetermined by the nature of your work, as with scientific reports. If that's the case, you can ignore this step and move on.
Research Report: A research report is a document prepared by an analyst or strategist who is a part of the investment research team in a stock brokerage or investment bank . A research report may ...
Note that field reports should be written in the past tense. With this in mind, most field reports in the social sciences include the following elements: I. Introduction The introduction should describe the research problem, the specific objectives of your research, and the important theories or concepts underpinning your field study.
Table of contents. Step 1: Restate the problem. Step 2: Sum up the paper. Step 3: Discuss the implications. Research paper conclusion examples. Frequently asked questions about research paper conclusions.
The initial 12 risk factors were linked with 40% of cases, but the new report suggests addressing the 14 risk factors could help eliminate or delay 45% of dementia cases, said Livingston, a ...
U.S. based short-seller Hindenburg Research alleged on Saturday that the head of India's market regulator, Madhabi Puri Buch, previously held investments in offshore funds also used by the Adani ...
Provide details only in the body of your report. So, this is the foundation on which you build the logical next step to reach a conclusion that answers your research question. Try to keep the structure of the introduction simple. An effective way is to start with a rather general statement about the topic.
This new research plan amplifies the school's ability to translate Stanford research into large-scale solutions and inform key decision makers in policy and business.
Short seller Hindenburg Research published a report alleging a conflict of interest by the head of India's market regulator and claimed this prevented a proper investigation into Adani.
Adani Group, the Indian conglomerate rocked by a Hindenburg Research report last year, faced another heavy share selloff on Monday after the shortseller accused the head of India's market ...
The CAFA challenge reports improved protein function prediction and new functional annotations for hundreds of genes through experimental screens. Genome Biol. 20 , 1-23 (2019).
Today, the Biden-Harris Administration announced that the U.S. Department of Commerce and SK hynix have signed a non-binding preliminary memorandum of terms (PMT) to provide up to $450 million in proposed federal incentives under the CHIPS and Science Act to establish a high-bandwidth memory (HBM) advanced packaging fabrication and research and ...